-
Preparing raw data
mkdir raw_data; cd raw_data # control samples (8) ln -s ../X101SC26025981-Z01-J001/01.RawData/1/1_1.fq.gz AYE-WT_ctr_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/1/1_2.fq.gz AYE-WT_ctr_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/2/2_1.fq.gz AYE-WT_ctr_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/2/2_2.fq.gz AYE-WT_ctr_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/3/3_1.fq.gz AYE-WT_ctr_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/3/3_2.fq.gz AYE-WT_ctr_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/4/4_1.fq.gz AYE-T_ctr_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/4/4_2.fq.gz AYE-T_ctr_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/5/5_1.fq.gz AYE-T_ctr_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/5/5_2.fq.gz AYE-T_ctr_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/6/6_1.fq.gz AYE-T_ctr_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/6/6_2.fq.gz AYE-T_ctr_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/7/7_1.fq.gz AYE-O_ctr_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/7/7_2.fq.gz AYE-O_ctr_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/8/8_1.fq.gz AYE-O_ctr_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/8/8_2.fq.gz AYE-O_ctr_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/9/9_1.fq.gz AYE-O_ctr_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/9/9_2.fq.gz AYE-O_ctr_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/10/10_1.fq.gz O-Trans_ctr_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/10/10_2.fq.gz O-Trans_ctr_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/11/11_1.fq.gz O-Trans_ctr_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/11/11_2.fq.gz O-Trans_ctr_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/12/12_1.fq.gz O-Trans_ctr_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/12/12_2.fq.gz O-Trans_ctr_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/1new/1new_1.fq.gz WT-Trans_ctr_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/1new/1new_2.fq.gz WT-Trans_ctr_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/2new/2new_1.fq.gz WT-Trans_ctr_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/2new/2new_2.fq.gz WT-Trans_ctr_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/3new/3new_1.fq.gz WT-Trans_ctr_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/3new/3new_2.fq.gz WT-Trans_ctr_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/49/49_1.fq.gz AYE-WT_ctr_solid_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/49/49_2.fq.gz AYE-WT_ctr_solid_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/50/50_1.fq.gz AYE-WT_ctr_solid_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/50/50_2.fq.gz AYE-WT_ctr_solid_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/51/51_1.fq.gz AYE-WT_ctr_solid_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/51/51_2.fq.gz AYE-WT_ctr_solid_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/52/52_1.fq.gz AYE-O_ctr_solid_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/52/52_2.fq.gz AYE-O_ctr_solid_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/53/53_1.fq.gz AYE-O_ctr_solid_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/53/53_2.fq.gz AYE-O_ctr_solid_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/54/54_1.fq.gz AYE-O_ctr_solid_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/54/54_2.fq.gz AYE-O_ctr_solid_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/55/55_1.fq.gz AYE-T_ctr_solid_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/55/55_2.fq.gz AYE-T_ctr_solid_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/56/56_1.fq.gz AYE-T_ctr_solid_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/56/56_2.fq.gz AYE-T_ctr_solid_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/57/57_1.fq.gz AYE-T_ctr_solid_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/57/57_2.fq.gz AYE-T_ctr_solid_r3_R2.fastq.gz # Diclofenac(双氯芬酸)treatment (6) ln -s ../X101SC26025981-Z01-J001/01.RawData/25/25_1.fq.gz AYE-WT_Diclo750_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/25/25_2.fq.gz AYE-WT_Diclo750_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/26/26_1.fq.gz AYE-WT_Diclo750_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/26/26_2.fq.gz AYE-WT_Diclo750_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/27/27_1.fq.gz AYE-WT_Diclo750_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/27/27_2.fq.gz AYE-WT_Diclo750_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/28/28_1.fq.gz AYE-T_Diclo375_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/28/28_2.fq.gz AYE-T_Diclo375_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/29/29_1.fq.gz AYE-T_Diclo375_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/29/29_2.fq.gz AYE-T_Diclo375_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/30/30_1.fq.gz AYE-T_Diclo375_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/30/30_2.fq.gz AYE-T_Diclo375_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/31/31_1.fq.gz AYE-O_Diclo375_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/31/31_2.fq.gz AYE-O_Diclo375_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/32/32_1.fq.gz AYE-O_Diclo375_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/32/32_2.fq.gz AYE-O_Diclo375_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/33/33_1.fq.gz AYE-O_Diclo375_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/33/33_2.fq.gz AYE-O_Diclo375_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/34/34_1.fq.gz O-Trans_Diclo375_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/34/34_2.fq.gz O-Trans_Diclo375_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/35/35_1.fq.gz O-Trans_Diclo375_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/35/35_2.fq.gz O-Trans_Diclo375_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/36/36_1.fq.gz O-Trans_Diclo375_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/36/36_2.fq.gz O-Trans_Diclo375_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/4new/4new_1.fq.gz WT-Trans_Diclo750_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/4new/4new_2.fq.gz WT-Trans_Diclo750_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/5new/5new_1.fq.gz WT-Trans_Diclo750_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/5new/5new_2.fq.gz WT-Trans_Diclo750_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/6new/6new_1.fq.gz WT-Trans_Diclo750_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/6new/6new_2.fq.gz WT-Trans_Diclo750_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/73/73_1.fq.gz AYE-WT_Diclo1250_solid_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/73/73_2.fq.gz AYE-WT_Diclo1250_solid_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/74/74_1.fq.gz AYE-WT_Diclo1250_solid_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/74/74_2.fq.gz AYE-WT_Diclo1250_solid_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/75/75_1.fq.gz AYE-WT_Diclo1250_solid_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/75/75_2.fq.gz AYE-WT_Diclo1250_solid_r3_R2.fastq.gz # Rifampicin(利福平)treatment (4) ln -s ../X101SC26025981-Z01-J001/01.RawData/13/13_1.fq.gz AYE-WT_Rifampicin1.5_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/13/13_2.fq.gz AYE-WT_Rifampicin1.5_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/14/14_1.fq.gz AYE-WT_Rifampicin1.5_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/14/14_2.fq.gz AYE-WT_Rifampicin1.5_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/15/15_1.fq.gz AYE-WT_Rifampicin1.5_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/15/15_2.fq.gz AYE-WT_Rifampicin1.5_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/16/16_1.fq.gz AYE-T_Rifampicin2_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/16/16_2.fq.gz AYE-T_Rifampicin2_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/17/17_1.fq.gz AYE-T_Rifampicin2_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/17/17_2.fq.gz AYE-T_Rifampicin2_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/18/18_1.fq.gz AYE-T_Rifampicin2_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/18/18_2.fq.gz AYE-T_Rifampicin2_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/19/19_1.fq.gz AYE-O_Rifampicin2_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/19/19_2.fq.gz AYE-O_Rifampicin2_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/20/20_1.fq.gz AYE-O_Rifampicin2_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/20/20_2.fq.gz AYE-O_Rifampicin2_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/21/21_1.fq.gz AYE-O_Rifampicin2_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/21/21_2.fq.gz AYE-O_Rifampicin2_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/22/22_1.fq.gz O-Trans_Rifampicin2_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/22/22_2.fq.gz O-Trans_Rifampicin2_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/23/23_1.fq.gz O-Trans_Rifampicin2_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/23/23_2.fq.gz O-Trans_Rifampicin2_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/24/24_1.fq.gz O-Trans_Rifampicin2_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/24/24_2.fq.gz O-Trans_Rifampicin2_r3_R2.fastq.gz # Meropenem(美罗培南)treatment (4) ln -s ../X101SC26025981-Z01-J001/01.RawData/37/37_1.fq.gz AYE-WT_Mero0.35-0.5_r1_R1.fastq.gz #AYE-WT_Mero0.5_r1 ln -s ../X101SC26025981-Z01-J001/01.RawData/37/37_2.fq.gz AYE-WT_Mero0.35-0.5_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/38/38_1.fq.gz AYE-WT_Mero0.35-0.5_r2_R1.fastq.gz #AYE-WT_YX_Mero0.35_r2 ln -s ../X101SC26025981-Z01-J001/01.RawData/38/38_2.fq.gz AYE-WT_Mero0.35-0.5_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/39/39_1.fq.gz AYE-WT_Mero0.35-0.5_r3_R1.fastq.gz #AYE-WT_public_Mero0.35_r3 ln -s ../X101SC26025981-Z01-J001/01.RawData/39/39_2.fq.gz AYE-WT_Mero0.35-0.5_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/40/40_1.fq.gz AYE-T_Mero0.15_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/40/40_2.fq.gz AYE-T_Mero0.15_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/41/41_1.fq.gz AYE-T_Mero0.15_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/41/41_2.fq.gz AYE-T_Mero0.15_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/42/42_1.fq.gz AYE-T_Mero0.15_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/42/42_2.fq.gz AYE-T_Mero0.15_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/43/43_1.fq.gz AYE-O_Mero0.5_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/43/43_2.fq.gz AYE-O_Mero0.5_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/44/44_1.fq.gz AYE-O_Mero0.5_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/44/44_2.fq.gz AYE-O_Mero0.5_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/45/45_1.fq.gz AYE-O_Mero0.5_r3_R1.fastq.gz #Mero0.45 ln -s ../X101SC26025981-Z01-J001/01.RawData/45/45_2.fq.gz AYE-O_Mero0.5_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/46/46_1.fq.gz O-Trans_Mero0.25_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/46/46_2.fq.gz O-Trans_Mero0.25_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/47/47_1.fq.gz O-Trans_Mero0.25_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/47/47_2.fq.gz O-Trans_Mero0.25_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/48/48_1.fq.gz O-Trans_Mero0.25_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/48/48_2.fq.gz O-Trans_Mero0.25_r3_R2.fastq.gz # Azithromycin(阿奇霉素)treatment (5), among them, F_ctr_solid is clinical isolate. ln -s ../X101SC26025981-Z01-J001/01.RawData/58/58_1.fq.gz F_ctr_solid_r1_R1.fastq.gz #clinical ln -s ../X101SC26025981-Z01-J001/01.RawData/58/58_2.fq.gz F_ctr_solid_r1_R2.fastq.gz #clinical ln -s ../X101SC26025981-Z01-J001/01.RawData/59/59_1.fq.gz F_ctr_solid_r2_R1.fastq.gz #clinical ln -s ../X101SC26025981-Z01-J001/01.RawData/59/59_2.fq.gz F_ctr_solid_r2_R2.fastq.gz #clinical ln -s ../X101SC26025981-Z01-J001/01.RawData/60/60_1.fq.gz F_ctr_solid_r3_R1.fastq.gz #clinical ln -s ../X101SC26025981-Z01-J001/01.RawData/60/60_2.fq.gz F_ctr_solid_r3_R2.fastq.gz #clinical ln -s ../X101SC26025981-Z01-J001/01.RawData/61/61_1.fq.gz AYE-WT_Azi20_solid_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/61/61_2.fq.gz AYE-WT_Azi20_solid_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/62/62_1.fq.gz AYE-WT_Azi20_solid_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/62/62_2.fq.gz AYE-WT_Azi20_solid_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/63/63_1.fq.gz AYE-WT_Azi20_solid_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/63/63_2.fq.gz AYE-WT_Azi20_solid_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/67/67_1.fq.gz AYE-T_Azi20_solid_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/67/67_2.fq.gz AYE-T_Azi20_solid_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/68/68_1.fq.gz AYE-T_Azi20_solid_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/68/68_2.fq.gz AYE-T_Azi20_solid_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/69/69_1.fq.gz AYE-T_Azi20_solid_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/69/69_2.fq.gz AYE-T_Azi20_solid_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/64/64_1.fq.gz AYE-O_Azi20_solid_r1_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/64/64_2.fq.gz AYE-O_Azi20_solid_r1_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/65/65_1.fq.gz AYE-O_Azi20_solid_r2_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/65/65_2.fq.gz AYE-O_Azi20_solid_r2_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/66/66_1.fq.gz AYE-O_Azi20_solid_r3_R1.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/66/66_2.fq.gz AYE-O_Azi20_solid_r3_R2.fastq.gz ln -s ../X101SC26025981-Z01-J001/01.RawData/70/70_1.fq.gz F_Azi20_solid_r1_R1.fastq.gz #clinical ln -s ../X101SC26025981-Z01-J001/01.RawData/70/70_2.fq.gz F_Azi20_solid_r1_R2.fastq.gz #clinical ln -s ../X101SC26025981-Z01-J001/01.RawData/71/71_1.fq.gz F_Azi20_solid_r2_R1.fastq.gz #clinical ln -s ../X101SC26025981-Z01-J001/01.RawData/71/71_2.fq.gz F_Azi20_solid_r2_R2.fastq.gz #clinical ln -s ../X101SC26025981-Z01-J001/01.RawData/72/72_1.fq.gz F_Azi20_solid_r3_R1.fastq.gz #clinical ln -s ../X101SC26025981-Z01-J001/01.RawData/72/72_2.fq.gz F_Azi20_solid_r3_R2.fastq.gz #clinical -
Preparing the directory trimmed
mkdir trimmed trimmed_unpaired; for sample_id in AYE-O_Azi20_solid_r1 AYE-O_Azi20_solid_r2 AYE-O_Azi20_solid_r3 AYE-O_ctr_r1 AYE-O_ctr_r2 AYE-O_ctr_r3 AYE-O_ctr_solid_r1 AYE-O_ctr_solid_r2 AYE-O_ctr_solid_r3 AYE-O_Diclo375_r1 AYE-O_Diclo375_r2 AYE-O_Diclo375_r3 AYE-O_Mero0.5_r1 AYE-O_Mero0.5_r2 AYE-O_Mero0.5_r3 AYE-O_Rifampicin2_r1 AYE-O_Rifampicin2_r2 AYE-O_Rifampicin2_r3 AYE-T_Azi20_solid_r1 AYE-T_Azi20_solid_r2 AYE-T_Azi20_solid_r3 AYE-T_ctr_r1 AYE-T_ctr_r2 AYE-T_ctr_r3 AYE-T_ctr_solid_r1 AYE-T_ctr_solid_r2 AYE-T_ctr_solid_r3 AYE-T_Diclo375_r1 AYE-T_Diclo375_r2 AYE-T_Diclo375_r3 AYE-T_Mero0.15_r1 AYE-T_Mero0.15_r2 AYE-T_Mero0.15_r3 AYE-T_Rifampicin2_r1 AYE-T_Rifampicin2_r2 AYE-T_Rifampicin2_r3 AYE-WT_Azi20_solid_r1 AYE-WT_Azi20_solid_r2 AYE-WT_Azi20_solid_r3 AYE-WT_ctr_r1 AYE-WT_ctr_r2 AYE-WT_ctr_r3 AYE-WT_ctr_solid_r1 AYE-WT_ctr_solid_r2 AYE-WT_ctr_solid_r3 AYE-WT_Diclo1250_solid_r1 AYE-WT_Diclo1250_solid_r2 AYE-WT_Diclo1250_solid_r3 AYE-WT_Diclo750_r1 AYE-WT_Diclo750_r2 AYE-WT_Diclo750_r3 AYE-WT_Mero0.35-0.5_r1 AYE-WT_Mero0.35-0.5_r2 AYE-WT_Mero0.35-0.5_r3 AYE-WT_Rifampicin1.5_r1 AYE-WT_Rifampicin1.5_r2 AYE-WT_Rifampicin1.5_r3 F_Azi20_solid_r1 F_Azi20_solid_r2 F_Azi20_solid_r3 F_ctr_solid_r1 F_ctr_solid_r2 F_ctr_solid_r3 O-Trans_ctr_r1 O-Trans_ctr_r2 O-Trans_ctr_r3 O-Trans_Diclo375_r1 O-Trans_Diclo375_r2 O-Trans_Diclo375_r3 O-Trans_Mero0.25_r1 O-Trans_Mero0.25_r2 O-Trans_Mero0.25_r3 O-Trans_Rifampicin2_r1 O-Trans_Rifampicin2_r2 O-Trans_Rifampicin2_r3 WT-Trans_ctr_r1 WT-Trans_ctr_r2 WT-Trans_ctr_r3 WT-Trans_Diclo750_r1 WT-Trans_Diclo750_r2 WT-Trans_Diclo750_r3; do \ for sample_id in AYE-T_Diclo375_r2; do \ java -jar /home/jhuang/Tools/Trimmomatic-0.36/trimmomatic-0.36.jar PE -threads 100 raw_data/${sample_id}_R1.fastq.gz raw_data/${sample_id}_R2.fastq.gz trimmed/${sample_id}_R1.fastq.gz trimmed_unpaired/${sample_id}_R1.fastq.gz trimmed/${sample_id}_R2.fastq.gz trimmed_unpaired/${sample_id}_R2.fastq.gz ILLUMINACLIP:/home/jhuang/Tools/Trimmomatic-0.36/adapters/TruSeq3-PE-2.fa:2:30:10:8:TRUE LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:36 AVGQUAL:20; done 2> trimmomatic_pe.log; done -
(Optional) using trinity to find the most closely reference
#Trinity --seqType fq --max_memory 50G --left trimmed/wt_r1_R1.fastq.gz --right trimmed/wt_r1_R2.fastq.gz --CPU 12 #https://www.genome.jp/kegg/tables/br08606.html#prok acb KGB Acinetobacter baumannii ATCC 17978 2007 GenBank abm KGB Acinetobacter baumannii SDF 2008 GenBank aby KGB Acinetobacter baumannii AYE 2008 GenBank --> * abc KGB Acinetobacter baumannii ACICU 2008 GenBank abn KGB Acinetobacter baumannii AB0057 2008 GenBank abb KGB Acinetobacter baumannii AB307-0294 2008 GenBank abx KGB Acinetobacter baumannii 1656-2 2012 GenBank abz KGB Acinetobacter baumannii MDR-ZJ06 2012 GenBank abr KGB Acinetobacter baumannii MDR-TJ 2012 GenBank abd KGB Acinetobacter baumannii TCDC-AB0715 2012 GenBank abh KGB Acinetobacter baumannii TYTH-1 2012 GenBank abad KGB Acinetobacter baumannii D1279779 2013 GenBank abj KGB Acinetobacter baumannii BJAB07104 2013 GenBank abab KGB Acinetobacter baumannii BJAB0715 2013 GenBank abaj KGB Acinetobacter baumannii BJAB0868 2013 GenBank abaz KGB Acinetobacter baumannii ZW85-1 2013 GenBank abk KGB Acinetobacter baumannii AbH12O-A2 2014 GenBank abau KGB Acinetobacter baumannii AB030 2014 GenBank abaa KGB Acinetobacter baumannii AB031 2014 GenBank abw KGB Acinetobacter baumannii AC29 2014 GenBank abal KGB Acinetobacter baumannii LAC-4 2015 GenBank #Note that the Acinetobacter baumannii strain ATCC 19606 chromosome, complete genome (GenBank: CU459141.1) was choosen as reference! -
Preparing samplesheet.csv
sample,fastq_1,fastq_2,strandedness Urine_r1,Urine_r1_R1.fq.gz,Urine_r1_R2.fq.gz,auto ... -
Downloading CU459141.fasta and CU459141.gff from GenBank and preparing CU459141_m.gff
#Example1: http://xgenes.com/article/article-content/157/prepare-virus-gtf-for-nextflow-run/ #Default NOT_WORKING: --gtf_group_features 'gene_id' --gtf_extra_attributes 'gene_name' --featurecounts_group_type 'gene_biotype' --featurecounts_feature_type 'exon' #(host_env) !NOT_WORKING! jhuang@WS-2290C:~/DATA/Data_Tam_RNAseq_2024$ /usr/local/bin/nextflow run rnaseq/main.nf --input samplesheet.csv --outdir results --fasta "/home/jhuang/DATA/Data_Tam_RNAseq_2024/CU459141.fasta" --gff "/home/jhuang/DATA/Data_Tam_RNAseq_2024/CU459141.gff" -profile docker -resume --max_cpus 55 --max_memory 512.GB --max_time 2400.h --save_align_intermeds --save_unaligned --save_reference --aligner 'star_salmon' --gtf_group_features 'gene_id' --gtf_extra_attributes 'gene_name' --featurecounts_group_type 'gene_biotype' --featurecounts_feature_type 'transcript' # -- DEBUG_1 (CDS --> exon in CP059040.gff) -- #Checking the record (see below) in results/genome/CP059040.gtf #In ./results/genome/CP059040.gtf e.g. "CP059040.1 Genbank transcript 1 1398 . + . transcript_id "gene-H0N29_00005"; gene_id "gene-H0N29_00005"; gene_name "dnaA"; Name "dnaA"; gbkey "Gene"; gene "dnaA"; gene_biotype "protein_coding"; locus_tag "H0N29_00005";" #--featurecounts_feature_type 'transcript' returns only the tRNA results #Since the tRNA records have "transcript and exon". In gene records, we have "transcript and CDS". replace the CDS with exon grep -P "\texon\t" CP059040.gff | sort | wc -l #96 grep -P "cmsearch\texon\t" CP059040.gff | wc -l #=10 ignal recognition particle sRNA small typ, transfer-messenger RNA, 5S ribosomal RNA grep -P "Genbank\texon\t" CP059040.gff | wc -l #=12 16S and 23S ribosomal RNA grep -P "tRNAscan-SE\texon\t" CP059040.gff | wc -l #tRNA 74 wc -l star_salmon/AUM_r3/quant.genes.sf #--featurecounts_feature_type 'transcript' results in 96 records! grep -P "\tCDS\t" CU459141.gff3 | wc -l #3659 sed 's/\tCDS\t/\texon\t/g' CU459141.gff3 > CU459141_m.gff grep -P "\texon\t" CU459141_m.gff | sort | wc -l #3760 # -- DEBUG_2: combination of 'CU459141_m.gff' and 'exon' results in ERROR, using 'transcript' instead! --gff "/home/jhuang/DATA/Data_Tam_RNAseq_2026_on_AYE/CU459141_m.gff" --featurecounts_feature_type 'transcript' # -- DEBUG_3: make sure the header of fasta is the same to the *_m.gff file -
nextflow run
# ---- SUCCESSFUL with directly downloaded gff3 and fasta from NCBI using docker after replacing 'CDS' with 'exon' ---- (host_env) mv trimmed/*.fastq.gz . (host_env) nextflow run nf-core/rnaseq -r 3.14.0 -profile docker \–input samplesheet.csv –outdir results –fasta “/home/jhuang/DATA/Data_Tam_RNAseq_2026_on_AYE/CU459141.fasta” –gff “/home/jhuang/DATA/Data_Tam_RNAseq_2026_on_AYE/CU459141_m.gff” -resume –max_cpus 90 –max_memory 900.GB –max_time 2400.h –save_align_intermeds –save_unaligned –save_reference –aligner ‘star_salmon’ –gtf_group_features ‘gene_id’ –gtf_extra_attributes ‘gene_name’ –featurecounts_group_type ‘gene_biotype’ –featurecounts_feature_type ‘transcript’
-
Import data and pca-plot
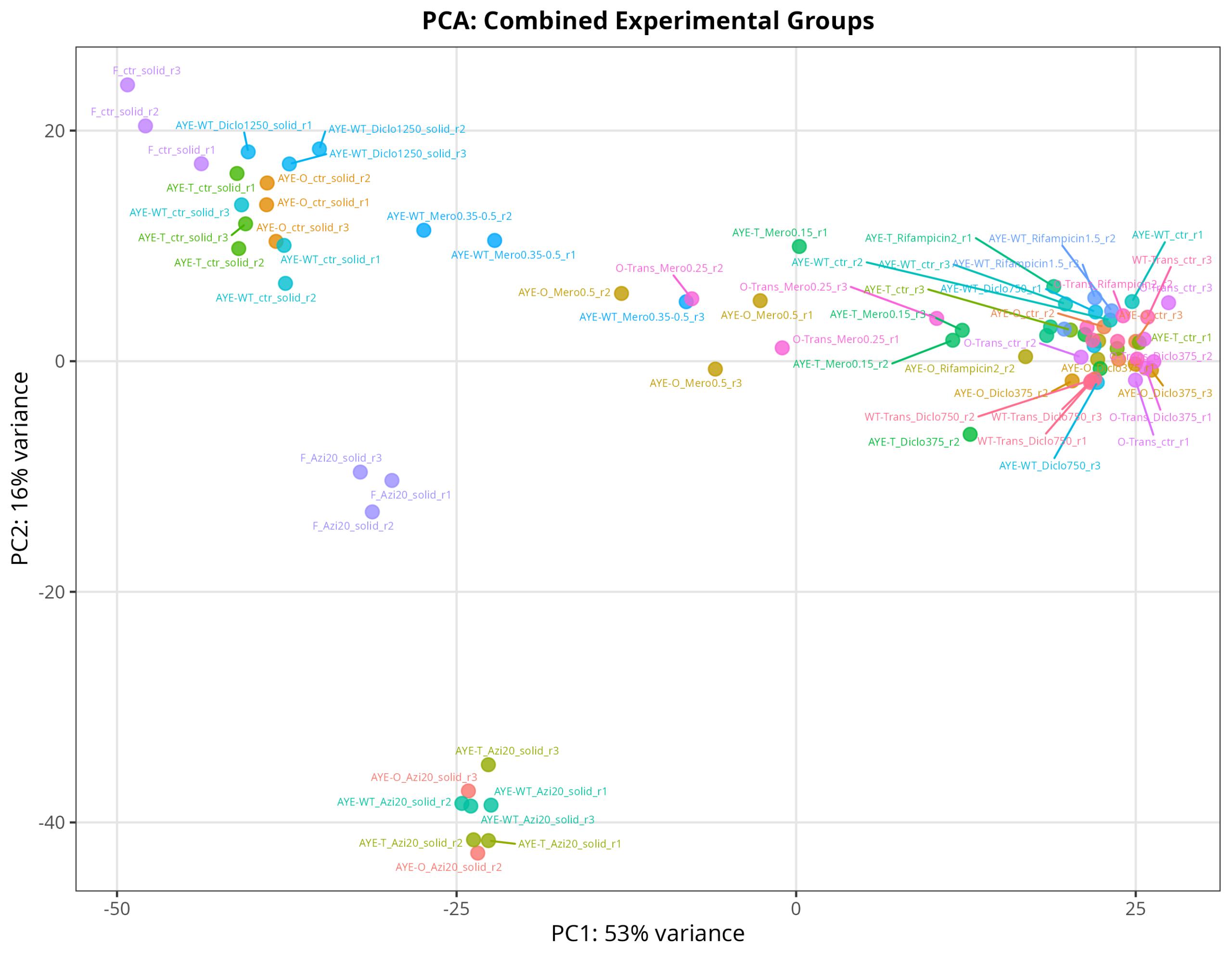
#TODO_AFTERNOON: at least sent today a PCA, am besten all comparison tables! #mamba activate r_env #install.packages("ggfun") # Import the required libraries library("AnnotationDbi") library("clusterProfiler") library("ReactomePA") library(gplots) library(tximport) library(DESeq2) #library("org.Hs.eg.db") library(dplyr) library(tidyverse) #install.packages("devtools") #devtools::install_version("gtable", version = "0.3.0") library(gplots) library("RColorBrewer") #install.packages("ggrepel") library("ggrepel") # install.packages("openxlsx") library(openxlsx) library(EnhancedVolcano) library(DESeq2) library(edgeR) setwd("~/DATA/Data_Tam_RNAseq_2026_on_AYE/results/star_salmon") # Define paths to your Salmon output quantification files # Store sample names in a character vector samples <- c( "AYE-O_Azi20_solid_r1", "AYE-O_Azi20_solid_r2", "AYE-O_Azi20_solid_r3", "AYE-O_ctr_r1", "AYE-O_ctr_r2", "AYE-O_ctr_r3", "AYE-O_ctr_solid_r1", "AYE-O_ctr_solid_r2", "AYE-O_ctr_solid_r3", "AYE-O_Diclo375_r1", "AYE-O_Diclo375_r2", "AYE-O_Diclo375_r3", "AYE-O_Mero0.5_r1", "AYE-O_Mero0.5_r2", "AYE-O_Mero0.5_r3", "AYE-O_Rifampicin2_r1", "AYE-O_Rifampicin2_r2", "AYE-O_Rifampicin2_r3", "AYE-T_Azi20_solid_r1", "AYE-T_Azi20_solid_r2", "AYE-T_Azi20_solid_r3", "AYE-T_ctr_r1", "AYE-T_ctr_r2", "AYE-T_ctr_r3", "AYE-T_ctr_solid_r1", "AYE-T_ctr_solid_r2", "AYE-T_ctr_solid_r3", "AYE-T_Diclo375_r1", "AYE-T_Diclo375_r2", "AYE-T_Diclo375_r3", "AYE-T_Mero0.15_r1", "AYE-T_Mero0.15_r2", "AYE-T_Mero0.15_r3", "AYE-T_Rifampicin2_r1", "AYE-T_Rifampicin2_r2", "AYE-T_Rifampicin2_r3", "AYE-WT_Azi20_solid_r1", "AYE-WT_Azi20_solid_r2", "AYE-WT_Azi20_solid_r3", "AYE-WT_ctr_r1", "AYE-WT_ctr_r2", "AYE-WT_ctr_r3", "AYE-WT_ctr_solid_r1", "AYE-WT_ctr_solid_r2", "AYE-WT_ctr_solid_r3", "AYE-WT_Diclo1250_solid_r1", "AYE-WT_Diclo1250_solid_r2", "AYE-WT_Diclo1250_solid_r3", "AYE-WT_Diclo750_r1", "AYE-WT_Diclo750_r2", "AYE-WT_Diclo750_r3", "AYE-WT_Mero0.35-0.5_r1", "AYE-WT_Mero0.35-0.5_r2", "AYE-WT_Mero0.35-0.5_r3", "AYE-WT_Rifampicin1.5_r1", "AYE-WT_Rifampicin1.5_r2", "AYE-WT_Rifampicin1.5_r3", "F_Azi20_solid_r1", "F_Azi20_solid_r2", "F_Azi20_solid_r3", "F_ctr_solid_r1", "F_ctr_solid_r2", "F_ctr_solid_r3", "O-Trans_ctr_r1", "O-Trans_ctr_r2", "O-Trans_ctr_r3", "O-Trans_Diclo375_r1", "O-Trans_Diclo375_r2", "O-Trans_Diclo375_r3", "O-Trans_Mero0.25_r1", "O-Trans_Mero0.25_r2", "O-Trans_Mero0.25_r3", "O-Trans_Rifampicin2_r1", "O-Trans_Rifampicin2_r2", "O-Trans_Rifampicin2_r3", "WT-Trans_ctr_r1", "WT-Trans_ctr_r2", "WT-Trans_ctr_r3", "WT-Trans_Diclo750_r1", "WT-Trans_Diclo750_r2", "WT-Trans_Diclo750_r3" ) ## Automatically generate the named vector files <- setNames(paste0("./", samples, "/quant.sf"), samples) # ----------------------------------------------------------------- # ---- Step 1: Create Detailed Metadata from Your Sample Names ---- # Extract metadata from sample names samples <- names(files) # Parse the complex sample names metadata <- data.frame( sample = samples, stringsAsFactors = FALSE ) # Extract strain (everything before first underscore or hyphen treatment) metadata$strain <- sapply(strsplit(samples, "[-_]"), function(x) { if(x[1] %in% c("AYE", "O", "WT", "F")) { if(x[1] == "AYE" && length(x) > 1 && x[2] %in% c("WT", "T", "O")) { paste(x[1:2], collapse = "-") } else if(x[1] %in% c("O", "WT") && x[2] == "Trans") { paste(x[1:2], collapse = "-") } else { x[1] } } else { x[1] } }) # Extract treatment type metadata$treatment <- sapply(samples, function(x) { if(grepl("_ctr", x)) return("ctrl") if(grepl("Diclo", x)) return("Diclo") if(grepl("Mero", x)) return("Mero") if(grepl("Azi", x)) return("Azi") if(grepl("Rifampicin", x)) return("Rifampicin") return("ctrl") }) # Extract concentration metadata$concentration <- sapply(samples, function(x) { if(grepl("Diclo1250", x)) return("1250") if(grepl("Diclo750", x)) return("750") if(grepl("Diclo375", x)) return("375") if(grepl("Mero0.5", x)) return("0.5") if(grepl("Mero0.35", x)) return("0.35") if(grepl("Mero0.25", x)) return("0.25") if(grepl("Mero0.15", x)) return("0.15") if(grepl("Azi20", x)) return("20") if(grepl("Rifampicin2", x)) return("2") if(grepl("Rifampicin1.5", x)) return("1.5") return("0") }) # Extract condition (solid vs liquid) metadata$condition <- ifelse(grepl("_solid", samples), "solid", "liquid") # Extract replicate metadata$replicate <- sapply(strsplit(samples, "_"), function(x) { rep_part <- x[length(x)] gsub("r", "", rep_part) }) # Create combined group for easy comparisons metadata$group <- paste(metadata$strain, metadata$treatment, metadata$concentration, sep = "_") # Set row names rownames(metadata) <- metadata$sample # Reorder to match txi columns metadata <- metadata[colnames(txi$counts), ] # --------------------------------------------- # ---- Step 2: Choose Your Design Strategy ---- # Strategy A: Full Factorial Design (if balanced) dds <- DESeqDataSetFromTximport(txi, metadata, design = ~ strain + treatment + condition) # --> Strategy B: Combined Group Factor ⭐ RECOMMENDED metadata$group <- factor(paste(metadata$strain, metadata$treatment, metadata$concentration, metadata$condition, sep = "_")) dds <- DESeqDataSetFromTximport(txi, metadata, design = ~ group) dds <- DESeq(dds) # See all available comparisons resultsNames(dds) # ------------------------------------------------------------- # ---- Step 3: Set Up Specific Comparisons from Your Notes ---- # ========================================== # 1. Define Exact Comparisons from Your Notes # ========================================== planned_comparisons <- list( # --- Baseline / Strain Controls --- AYE_T_ctr_vs_AYE_WT_ctr = list(treat = "AYE-T_ctrl_0_liquid", ctrl = "AYE-WT_ctrl_0_liquid"), AYE_O_ctr_vs_AYE_WT_ctr = list(treat = "AYE-O_ctrl_0_liquid", ctrl = "AYE-WT_ctrl_0_liquid"), O_Trans_ctr_vs_AYE_WT_ctr = list(treat = "O-Trans_ctrl_0_liquid", ctrl = "AYE-WT_ctrl_0_liquid"), WT_Trans_ctr_vs_AYE_WT_ctr = list(treat = "WT-Trans_ctrl_0_liquid",ctrl = "AYE-WT_ctrl_0_liquid"), AYE_O_ctr_vs_AYE_T = list(treat = "AYE-O_ctrl_0_liquid", ctrl = "AYE-T_ctrl_0_liquid"), O_Trans_ctr_vs_AYE_T = list(treat = "O-Trans_ctrl_0_liquid", ctrl = "AYE-T_ctrl_0_liquid"), WT_Trans_ctr_vs_AYE_T = list(treat = "WT-Trans_ctrl_0_liquid",ctrl = "AYE-T_ctrl_0_liquid"), # --- Condition Effects (Solid vs Liquid) --- AYE_WT_ctr_solid_vs_AYE_WT_ctr = list(treat = "AYE-WT_ctrl_0_solid", ctrl = "AYE-WT_ctrl_0_liquid"), AYE_O_ctr_solid_vs_AYE_O_ctr = list(treat = "AYE-O_ctrl_0_solid", ctrl = "AYE-O_ctrl_0_liquid"), AYE_T_ctr_solid_vs_AYE_T_ctr = list(treat = "AYE-T_ctrl_0_solid", ctrl = "AYE-T_ctrl_0_liquid"), AYE_O_ctr_solid_vs_AYE_WT_ctr_solid= list(treat = "AYE-O_ctrl_0_solid", ctrl = "AYE-WT_ctrl_0_solid"), AYE_T_ctr_solid_vs_AYE_WT_ctr_solid= list(treat = "AYE-T_ctrl_0_solid", ctrl = "AYE-WT_ctrl_0_solid"), # --- Diclofenac --- AYE_WT_Diclo750_vs_AYE_WT_ctr = list(treat = "AYE-WT_Diclo_750_liquid", ctrl = "AYE-WT_ctrl_0_liquid"), AYE_T_Diclo375_vs_AYE_WT_ctr = list(treat = "AYE-T_Diclo_375_liquid", ctrl = "AYE-WT_ctrl_0_liquid"), AYE_O_Diclo375_vs_AYE_WT_ctr = list(treat = "AYE-O_Diclo_375_liquid", ctrl = "AYE-WT_ctrl_0_liquid"), O_Trans_Diclo375_vs_AYE_WT_ctr = list(treat = "O-Trans_Diclo_375_liquid", ctrl = "AYE-WT_ctrl_0_liquid"), WT_Trans_Diclo750_vs_AYE_WT_ctr = list(treat = "WT-Trans_Diclo_750_liquid", ctrl = "AYE-WT_ctrl_0_liquid"), Diclo_AYE_WT_1250_solid_vs_solid_ctr = list(treat = "AYE-WT_Diclo_1250_solid", ctrl = "AYE-WT_ctrl_0_solid"), # --- Meropenem --- AYE_WT_Mero_vs_AYE_WT_ctr = list(treat = "AYE-WT_Mero_0.35_liquid", ctrl = "AYE-WT_ctrl_0_liquid"), AYE_T_Mero_vs_AYE_WT_ctr = list(treat = "AYE-T_Mero_0.15_liquid", ctrl = "AYE-WT_ctrl_0_liquid"), AYE_O_Mero_vs_AYE_WT_ctr = list(treat = "AYE-O_Mero_0.5_liquid", ctrl = "AYE-WT_ctrl_0_liquid"), O_Trans_Mero_vs_AYE_WT_ctr = list(treat = "O-Trans_Mero_0.25_liquid", ctrl = "AYE-WT_ctrl_0_liquid"), AYE_T_Mero_vs_AYE_T_ctr = list(treat = "AYE-T_Mero_0.15_liquid", ctrl = "AYE-T_ctrl_0_liquid"), # --- Azithromycin (Solid) --- AYE_WT_Azi_vs_solid_ctr = list(treat = "AYE-WT_Azi_20_solid", ctrl = "AYE-WT_ctrl_0_solid"), AYE_T_Azi_vs_solid_ctr = list(treat = "AYE-T_Azi_20_solid", ctrl = "AYE-T_ctrl_0_solid"), AYE_O_Azi_vs_solid_ctr = list(treat = "AYE-O_Azi_20_solid", ctrl = "AYE-O_ctrl_0_solid"), F_Azi_vs_F_solid_ctr = list(treat = "F_Azi_20_solid", ctrl = "F_ctrl_0_solid"), # --- Rifampicin --- AYE_WT_Rif_vs_AYE_WT_ctr = list(treat = "AYE-WT_Rifampicin_1.5_liquid", ctrl = "AYE-WT_ctrl_0_liquid"), AYE_T_Rif_vs_AYE_T_ctr = list(treat = "AYE-T_Rifampicin_2_liquid", ctrl = "AYE-T_ctrl_0_liquid"), AYE_O_Rif_vs_AYE_O_ctr = list(treat = "AYE-O_Rifampicin_2_liquid", ctrl = "AYE-O_ctrl_0_liquid"), O_Trans_Rif_vs_O_Trans_ctr = list(treat = "O-Trans_Rifampicin_2_liquid", ctrl = "O-Trans_ctrl_0_liquid") ) # ========================================== # 2. Verification & Validation Script # ========================================== # Identify which column in colData holds your group names group_col <- if("group" %in% colnames(colData(dds))) "group" else if("treatment" %in% colnames(colData(dds))) "treatment" else stop("❌ Please specify the correct colData column containing group names.") actual_groups <- unique(colData(dds)[[group_col]]) cat("\n", paste(rep("=", 85), collapse=""), "\n") cat("📋 VERIFICATION OF NOTE-DERIVED COMPARISONS\n") cat(paste(rep("=", 85), collapse=""), "\n\n") validation_results <- data.frame( Comparison_Name = character(), Treatment_String = character(), Control_String = character(), Status = character(), Suggested_Contrast = character(), stringsAsFactors = FALSE ) for(name in names(planned_comparisons)) { trt <- planned_comparisons[[name]]$treat ctl <- planned_comparisons[[name]]$ctrl # Find closest matches in actual data trt_match <- actual_groups[grepl(trt, actual_groups, fixed = TRUE)] ctl_match <- actual_groups[grepl(ctl, actual_groups, fixed = TRUE)] status <- if(length(trt_match) > 0 && length(ctl_match) > 0) "✅ VALID" else "⚠️ CHECK" contrast_str <- if(status == "✅ VALID") paste0('c("', group_col, '", "', trt_match[1], '", "', ctl_match[1], '")') else "N/A" validation_results <- rbind(validation_results, data.frame( Comparison_Name = name, Treatment_String = trt, Control_String = ctl, Status = status, Suggested_Contrast = contrast_str, stringsAsFactors = FALSE )) cat(sprintf("%-45s | T:%-25s C:%-20s | %s\n", name, trt, ctl, status)) if(status == "⚠️ CHECK") { if(length(trt_match) == 0) cat(" 🔍 Treat not found. Closest: ", paste(head(actual_groups[grepl(strsplit(trt, "_")[[1]][1], actual_groups)], 3), collapse=", "), "\n") if(length(ctl_match) == 0) cat(" 🔍 Ctrl not found. Closest: ", paste(head(actual_groups[grepl(strsplit(ctl, "_")[[1]][1], actual_groups)], 3), collapse=", "), "\n") } } # ========================================== # 3. Auto-Generate DESeq2 results() Calls (Optional) # ========================================== valid_comparisons <- validation_results[validation_results$Status == "✅ VALID", ] if(nrow(valid_comparisons) > 0) { cat("\n📜 READY-TO-RUN DESeq2 CONTRASTS:\n") cat(paste(rep("-", 60), collapse=""), "\n") for(i in seq_len(nrow(valid_comparisons))) { cat(sprintf('res_%s <- results(dds, contrast = %s)\n', gsub("[^A-Za-z0-9]", "_", valid_comparisons$Comparison_Name[i]), valid_comparisons$Suggested_Contrast[i])) } } else { cat("\n⚠️ No exact matches found. Check your colData group naming convention.\n") } # ----------------------------- # ---- Step 4: PCA figures ---- # 🔍 What each figure shows: # # 01_PCA_by_Strain.png → Tests if genetic background (AYE-WT, AYE-T, AYE-O, Trans, F) is the dominant source of variation. # 02_PCA_by_Treatment.png → Shows clustering by antibiotic/drug exposure (ctrl, Diclo, Mero, Azi, Rifampicin). # 03_PCA_by_Condition.png → Reveals batch/growth media effects (solid vs liquid). # 04_PCA_CombinedGroups.png → Full experimental grouping with labeled sample names for quick outlier detection. # 05_PCA_Ellipses.png → Adds 95% confidence boundaries per strain to visualize group spread and overlap. # # ⚠️ Quick Checklist Before Running: # # Ensure metadata columns (strain, treatment, condition, group) are attached to colData(dds). # If ggrepel is missing, run install.packages("ggrepel"). # All PNGs will save to your current working directory (getwd()). # Install if missing: install.packages(c("ggplot2", "ggrepel")) library(DESeq2) library(ggplot2) library(ggrepel) # 1. Variance Stabilizing Transformation & Extract PCA Data vsd <- vst(dds, blind = FALSE) pca_data <- plotPCA(vsd, intgroup = c("strain", "treatment", "condition", "group"), returnData = TRUE) percent_var <- round(100 * attr(pca_data, "percentVar")) # Consistent theme for all plots base_theme <- theme_bw(base_size = 12) + theme(plot.title = element_text(hjust = 0.5, face = "bold", size = 13), legend.position = "right", legend.title = element_text(face = "bold"), panel.grid.major = element_line(color = "grey90"), panel.grid.minor = element_blank()) # --- Plot 1: Colored by Strain --- p1 <- ggplot(pca_data, aes(x = PC1, y = PC2, color = strain, shape = condition)) + geom_point(size = 3, alpha = 0.8) + geom_text_repel(aes(label = name), size = 2.5, max.overlaps = 20, show.legend = FALSE) + labs(x = paste0("PC1: ", percent_var[1], "% variance"), y = paste0("PC2: ", percent_var[2], "% variance"), title = "PCA: Samples Colored by Strain", color = "Strain", shape = "Condition") + base_theme ggsave("01_PCA_by_Strain.png", p1, width = 8, height = 6, dpi = 300) # --- Plot 2: Colored by Treatment --- p2 <- ggplot(pca_data, aes(x = PC1, y = PC2, color = treatment, shape = condition)) + geom_point(size = 3, alpha = 0.8) + labs(x = paste0("PC1: ", percent_var[1], "% variance"), y = paste0("PC2: ", percent_var[2], "% variance"), title = "PCA: Samples Colored by Treatment", color = "Treatment", shape = "Condition") + base_theme ggsave("02_PCA_by_Treatment.png", p2, width = 8, height = 6, dpi = 300) # --- Plot 3: Colored by Condition (Solid vs Liquid) --- p3 <- ggplot(pca_data, aes(x = PC1, y = PC2, color = condition, shape = strain)) + geom_point(size = 3, alpha = 0.8) + labs(x = paste0("PC1: ", percent_var[1], "% variance"), y = paste0("PC2: ", percent_var[2], "% variance"), title = "PCA: Samples Colored by Growth Condition", color = "Condition", shape = "Strain") + base_theme ggsave("03_PCA_by_Condition.png", p3, width = 8, height = 6, dpi = 300) # --- Plot 4: Combined Groups with Sample Labels --- p4 <- ggplot(pca_data, aes(x = PC1, y = PC2, color = group)) + geom_point(size = 3, alpha = 0.8) + geom_text_repel(aes(label = name), size = 2, max.overlaps = 30, box.padding = 0.3) + labs(x = paste0("PC1: ", percent_var[1], "% variance"), y = paste0("PC2: ", percent_var[2], "% variance"), title = "PCA: Combined Experimental Groups", color = "Group") + base_theme + theme(legend.position = "none") ggsave("04_PCA_CombinedGroups.png", p4, width = 9, height = 7, dpi = 300) # --- Plot 5: 95% Confidence Ellipses (by Strain) --- p5 <- ggplot(pca_data, aes(x = PC1, y = PC2, color = strain, fill = strain)) + geom_point(size = 3, alpha = 0.7) + stat_ellipse(level = 0.95, alpha = 0.2, geom = "polygon", show.legend = FALSE) + labs(x = paste0("PC1: ", percent_var[1], "% variance"), y = paste0("PC2: ", percent_var[2], "% variance"), title = "PCA: 95% Confidence Ellipses by Strain", color = "Strain", fill = "Strain") + base_theme ggsave("05_PCA_Ellipses.png", p5, width = 8, height = 6, dpi = 300) message("✅ All 5 PCA plots saved to working directory!") -
Run Differential Expression & PCA Analysis Complete
(r_env) jhuang@WS-2290C:/mnt/md1/DATA/Data_Tam_RNAseq_2026_on_AYE/results/star_salmon$ Rscript complete_deg_pipeline.R
RNAseq Data_Tam_RNAseq_2026_on_AYE using Rscript
Leave a reply