Microbiome analysis
Sample collection and storage
Fecal samples were collected from mice between 9–10 am at the indicated timepoints and immediately frozen at −80 °C until use.
DNA extraction
Genomic DNA was extracted from fecal pellets using the QIAamp Fast DNA Stool Mini Kit (Qiagen, #51604) following a protocol optimized for murine stool. Briefly, pellets were mechanically homogenized in a Precellys® 24 homogenizer (Bertin Technologies) using Soil grinding tubes (Bertin Technologies, #SK38), and DNA was extracted according to the manufacturer’s instructions. DNA yield was measured with NanoDrop, and the concentration was adjusted to 10 ng/µl.
16S rRNA gene amplicon library preparation and sequencing
16S rRNA amplicons (V3–V4 region) were generated using degenerate primers that contain Illumina adapter overhang sequences: forward (5′-TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG-3′) and reverse (5′-GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC-3′), following established Illumina 16S library preparation guidance. [1] Samples were multiplexed using the Nextera XT Index Kit (Illumina, #FC-131-1001) and sequenced by 500-cycle paired-end sequencing (2×250 bp) on an Illumina MiSeq (Illumina, #MS-102-2003).
Bioinformatics processing and QC
Raw FASTQ files were inspected with FastQC. [2] Sequence processing was performed in QIIME 2. [3] Demultiplexed paired-end reads were imported using a Phred33 manifest and per-base quality profiles were reviewed to guide trimming/truncation. ASVs were inferred using the QIIME 2 DADA2 workflow. [4] Primer/adapter-proximal bases were removed using fixed 5′ trimming (17 bp forward; 21 bp reverse), reads were filtered using expected-error thresholds, and multiple forward/reverse truncation length combinations were evaluated to optimize read retention and paired-end merging for the ~460 bp V3–V4 amplicon. Final denoising parameters were selected based on denoising statistics (successful merging and proportion of non-chimeric reads across samples), and the resulting feature table and representative ASV sequences were used for downstream analyses.
Taxonomy was assigned by VSEARCH-based consensus classification [5] against SILVA (release 132) [6] at 97% identity. Downstream visualization and statistics were performed in R using phyloseq. [7]
Diversity analyses, ordination, and statistics
Alpha diversity (Observed features/OTU richness, Shannon diversity, and Faith’s phylogenetic diversity) was calculated on rarefied data and compared between groups using nonparametric rank-based tests (Wilcoxon rank-sum for two-group comparisons; Kruskal–Wallis for ≥3 groups with post-hoc pairwise Wilcoxon tests). [8–10]
Group differences in community composition (beta-diversity) were assessed using Bray–Curtis dissimilarities [11] (computed after Hellinger transformation of abundance profiles) [12] and tested using PERMANOVA (adonis2 in vegan) with permutation-based significance. [13–14] Ordination was performed by principal coordinates analysis (PCoA) on the Bray–Curtis distance matrix to visualize sample-to-sample relationships. [15]
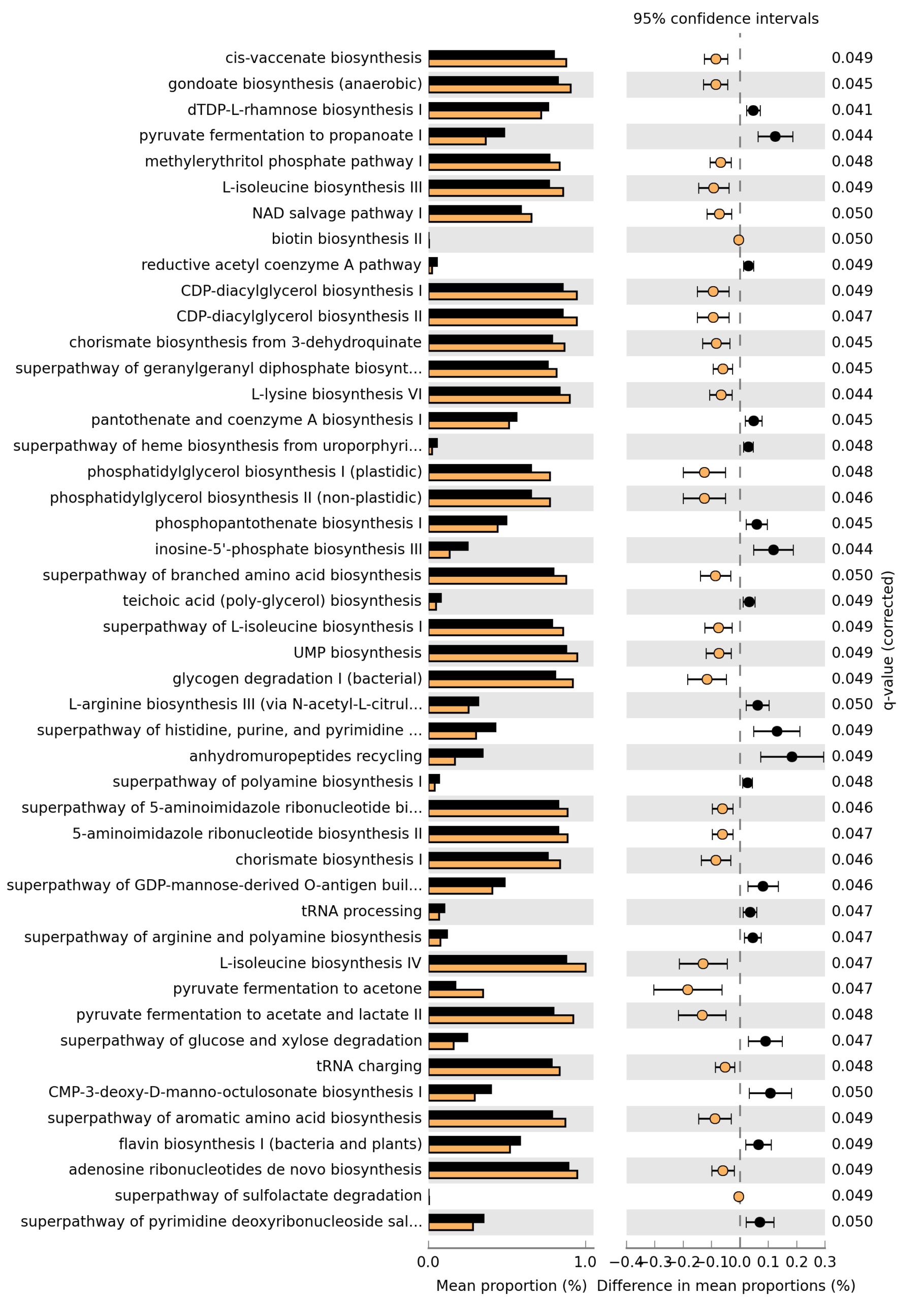
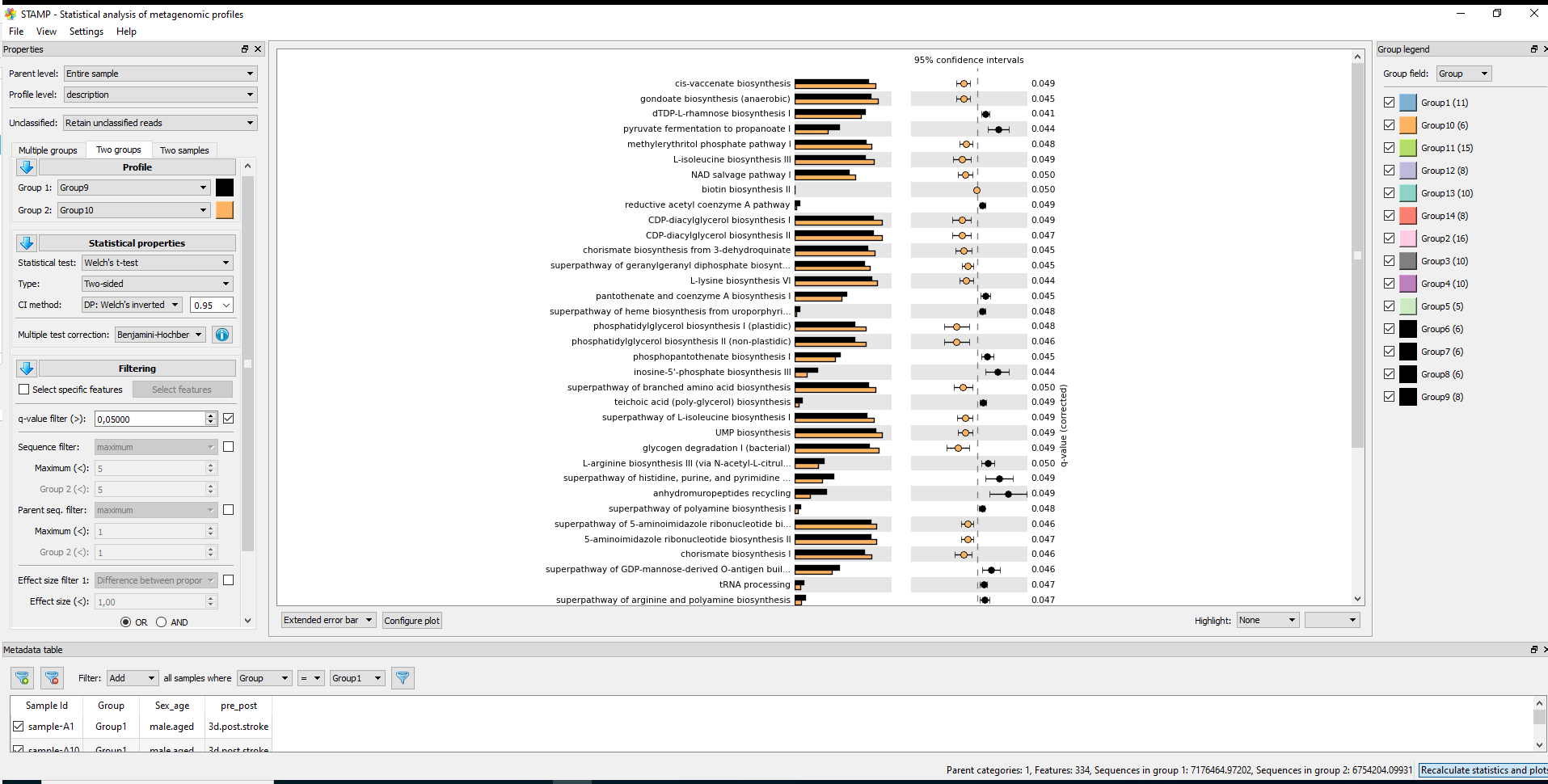
Differential abundance testing was performed using DESeq2 (negative binomial GLM). [16] Across analyses involving multiple hypothesis tests, p-values were adjusted using the Benjamini–Hochberg false discovery rate procedure. [17]
Pathway inference
Predicted functional potential (gene families and pathways) was inferred from 16S ASVs using PICRUSt2. [18]
References
Illumina. 16S Metagenomic Sequencing Library Preparation (Part #15044223 Rev. B) [Internet]. Illumina; 2013. (Includes the overhang adapter primer structure and V3–V4 primer sequences).
Andrews S. FastQC: a quality control tool for high throughput sequence data [Internet]. Babraham Bioinformatics; 2010 [cited 2025 Dec 18]. Available from: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol. 2019;37(8):852–857. doi:10.1038/s41587-019-0209-9.
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP. DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods. 2016;13(7):581–583. doi:10.1038/nmeth.3869.
Rognes T, Flouri T, Nichols B, Quince C, Mahé F. VSEARCH: a versatile open source tool for metagenomics. PeerJ. 2016;4:e2584. doi:10.7717/peerj.2584.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41(D1):D590–D596. doi:10.1093/nar/gks1219.
McMurdie PJ, Holmes S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One. 2013;8(4):e61217. doi:10.1371/journal.pone.0061217.
Wilcoxon F. Individual comparisons by ranking methods. Biometrics Bulletin. 1945;1(6):80–83. doi:10.2307/3001968.
Kruskal WH, Wallis WA. Use of ranks in one-criterion variance analysis. J Am Stat Assoc. 1952;47:583–621. doi:10.1080/01621459.1952.10483441.
(If you want an additional general nonparametric/post-hoc reference beyond Wilcoxon/Kruskal–Wallis, tell me your preferred one and I’ll format it consistently.)
Bray JR, Curtis JT. An ordination of the upland forest communities of southern Wisconsin. Ecol Monogr. 1957;27:325–349. doi:10.2307/1942268.
Legendre P, Gallagher ED. Ecologically meaningful transformations for ordination of species data. Oecologia. 2001;129(2):271–280. doi:10.1007/s004420100716.
Anderson MJ. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 2001;26:32–46. doi:10.1111/j.1442-9993.2001.01070.pp.x.
Oksanen J, Simpson GL, Blanchet FG, Kindt R, Legendre P, Minchin PR, et al. vegan: Community Ecology Package [Internet]. R package. Available from: https://CRAN.R-project.org/package=vegan. doi:10.32614/CRAN.package.vegan.
Gower JC. Some distance properties of latent root and vector methods used in multivariate analysis. Biometrika. 1966;53(3–4):325–338. doi:10.1093/biomet/53.3-4.325.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550. doi:10.1186/s13059-014-0550-8.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Series B. 1995;57(1):289–300. doi:10.1111/j.2517-6161.1995.tb02031.x.
Douglas GM, Maffei VJ, Zaneveld JR, Yurgel SN, Brown JR, Taylor CM, et al. PICRUSt2 for prediction of metagenome functions. Nat Biotechnol. 2020;38(6):685–688. doi:10.1038/s41587-020-0548-6. (slamo.biochem.dal.ca)
Phyloseq_Group9_10_11_pre-FMT.Rmd
author: “”
date: ‘r format(Sys.time(), "%d %m %Y")‘
header-includes:
-
\usepackage{color, fancyvrb}
output:
rmdformats::readthedown:
highlight: kate
number_sections : yes
pdf_document:
toc: yes
toc_depth: 2
number_sections : yes
#install.packages(c("picante", "rmdformats"))
#mamba install -c conda-forge freetype libpng harfbuzz fribidi
#mamba install -c conda-forge r-systemfonts r-svglite r-kableExtra freetype fontconfig harfbuzz fribidi libpng
library(knitr)
library(rmdformats)
library(readxl)
library(dplyr)
library(kableExtra)
library(openxlsx)
library(DESeq2)
library(writexl)
options(max.print="75")
knitr::opts_chunk$set(fig.width=8,
fig.height=6,
eval=TRUE,
cache=TRUE,
echo=TRUE,
prompt=FALSE,
tidy=FALSE,
comment=NA,
message=FALSE,
warning=FALSE)
opts_knit$set(width=85)
#rmarkdown::render('Phyloseq_Group9_10_11_pre-FMT.Rmd',output_file='Phyloseq_Group9_10_11_pre-FMT.html')
# Phyloseq R library
#* Phyloseq web site : https://joey711.github.io/phyloseq/index.html
#* See in particular tutorials for
# - importing data: https://joey711.github.io/phyloseq/import-data.html
# - heat maps: https://joey711.github.io/phyloseq/plot_heatmap-examples.html
Data
Import raw data and assign sample key:
#extend qiime2_metadata_for_qza_to_phyloseq.tsv with Diet and Flora
#setwd("~/DATA/Data_Laura_16S_2/core_diversity_e4753")
#map_corrected <- read.csv("qiime2_metadata_for_qza_to_phyloseq.tsv", sep="\t", row.names=1)
#knitr::kable(map_corrected) %>% kable_styling(bootstrap_options = c("striped", "hover", "condensed", "responsive"))
Prerequisites to be installed
install.packages("dplyr") # To manipulate dataframes
install.packages("readxl") # To read Excel files into R
install.packages("ggplot2") # for high quality graphics
install.packages("heatmaply")
source("https://bioconductor.org/biocLite.R")
biocLite("phyloseq")
#mamba install -c conda-forge r-ggplot2 r-vegan r-data.table
#BiocManager::install("microbiome")
#install.packages("ggpubr")
#install.packages("heatmaply")
library("readxl") # necessary to import the data from Excel file
library("ggplot2") # graphics
library("picante")
library("microbiome") # data analysis and visualisation
library("phyloseq") # also the basis of data object. Data analysis and visualisation
library("ggpubr") # publication quality figures, based on ggplot2
library("dplyr") # data handling, filter and reformat data frames
library("RColorBrewer") # nice color options
library("heatmaply")
library(vegan)
library(gplots)
#install.packages("openxlsx")
library(openxlsx)
Read the data and create phyloseq objects
Three tables are needed
library(tidyr)
# For QIIME1
#ps.ng.tax <- import_biom("./exported_table/feature-table.biom", "./exported-tree/tree.nwk")
# For QIIME2
#install.packages("remotes")
#remotes::install_github("jbisanz/qiime2R")
#"core_metrics_results/rarefied_table.qza", rarefying performed in the code, therefore import the raw table.
library(qiime2R)
ps_raw <- qza_to_phyloseq(
features = "table.qza", #cp ../Data_Karoline_16S_2025/dada2_tests2/test_7_f240_r240/table.qza .
tree = "rooted-tree.qza", #cp ../Data_Karoline_16S_2025/rooted-tree.qza .
metadata = "qiime2_metadata_for_qza_to_phyloseq.tsv" #cp ../Data_Karoline_16S_2025/qiime2_metadata_for_qza_to_phyloseq.tsv .
)
# or
#biom convert \
# -i ./exported_table/feature-table.biom \
# -o ./exported_table/feature-table-v1.biom \
# --to-json
#ps_raw <- import_biom("./exported_table/feature-table-v1.biom", treefilename="./exported-tree/tree.nwk")
sample <- read.csv("./qiime2_metadata_for_qza_to_phyloseq.tsv", sep="\t", row.names=1)
SAM = sample_data(sample, errorIfNULL = T)
#> setdiff(rownames(SAM), sample_names(ps_raw))
#[1] "sample-L9" should be removed since the low reads
ps_base <- merge_phyloseq(ps_raw, SAM)
print(ps_base)
taxonomy <- read.delim("taxonomy.tsv", sep="\t", header=TRUE) #cp ../Data_Karoline_16S_2025/exported-taxonomy/taxonomy.tsv .
#head(taxonomy)
# Separate taxonomy string into separate ranks
taxonomy_df <- taxonomy %>% separate(Taxon, into = c("Domain","Phylum","Class","Order","Family","Genus","Species"), sep = ";", fill = "right", extra = "drop")
# Use Feature.ID as rownames
rownames(taxonomy_df) <- taxonomy_df$Feature.ID
taxonomy_df <- taxonomy_df[, -c(1, ncol(taxonomy_df))] # Drop Feature.ID and Confidence
# Create tax_table
tax_table_final <- phyloseq::tax_table(as.matrix(taxonomy_df))
# Merge tax_table with existing phyloseq object
ps_base <- merge_phyloseq(ps_base, tax_table_final)
# Check
ps_base
#colnames(phyloseq::tax_table(ps_base)) <- c("Domain","Phylum","Class","Order","Family","Genus","Species")
saveRDS(ps_base, "./ps_base.rds")
Visualize data
sample_names(ps_base)
rank_names(ps_base)
sample_variables(ps_base)
# Define sample names once
samples <- c(
#"sample-A1","sample-A2","sample-A5","sample-A6","sample-A7","sample-A8","sample-A9","sample-A10", #RESIZED: "sample-A3","sample-A4","sample-A11",
#"sample-B1","sample-B2","sample-B3","sample-B4","sample-B5","sample-B6","sample-B7", #RESIZED: "sample-B8","sample-B9","sample-B10","sample-B11","sample-B12","sample-B13","sample-B14","sample-B15","sample-B16",
#"sample-C1","sample-C2","sample-C3","sample-C4","sample-C5","sample-C6","sample-C7", #RESIZED: "sample-C8","sample-C9","sample-C10",
#"sample-E1","sample-E2","sample-E3","sample-E4","sample-E5","sample-E6","sample-E7","sample-E8","sample-E9","sample-E10", #RESIZED:
#"sample-F1","sample-F2","sample-F3","sample-F4","sample-F5",
"sample-G1","sample-G2","sample-G3","sample-G4","sample-G5","sample-G6",
"sample-H1","sample-H2","sample-H3","sample-H4","sample-H5","sample-H6",
"sample-I1","sample-I2","sample-I3","sample-I4","sample-I5","sample-I6",
"sample-J1","sample-J2","sample-J3","sample-J4","sample-J10","sample-J11", #RESIZED: "sample-J5","sample-J8","sample-J9", "sample-J6","sample-J7",
"sample-K1","sample-K2","sample-K3","sample-K4","sample-K5","sample-K6", #RESIZED: "sample-K7","sample-K8","sample-K9","sample-K10", "sample-K11","sample-K12","sample-K13","sample-K14","sample-K15",
"sample-L2","sample-L3","sample-L4","sample-L5","sample-L6" #RESIZED:"sample-L1","sample-L7","sample-L8","sample-L10","sample-L11","sample-L12","sample-L13","sample-L14","sample-L15",
#"sample-M1","sample-M2","sample-M3","sample-M4","sample-M5","sample-M6","sample-M7","sample-M8",
#"sample-N1","sample-N2","sample-N3","sample-N4","sample-N5","sample-N6","sample-N7","sample-N8","sample-N9","sample-N10",
#"sample-O1","sample-O2","sample-O3","sample-O4","sample-O5","sample-O6","sample-O7","sample-O8"
)
ps_pruned <- prune_samples(samples, ps_base)
sample_names(ps_pruned)
rank_names(ps_pruned)
sample_variables(ps_pruned)
No samples were excluded as low-depth outliers (library sizes below the minimum depth threshold of 1,000 reads), and the remaining dataset (ps_filt) contains only samples meeting this depth cutoff with taxa retained only if they have nonzero total counts.
# ------------------------------------------------------------
# Filter low-depth samples (recommended for all analyses)
# ------------------------------------------------------------
min_depth <- 1000 # <-- adjust to your data / study design, keeps all!
ps_filt <- prune_samples(sample_sums(ps_pruned) >= min_depth, ps_pruned)
ps_filt <- prune_taxa(taxa_sums(ps_filt) > 0, ps_filt)
# Keep a depth summary for reporting / QC
depth_summary <- summary(sample_sums(ps_filt))
depth_summary
Differential abundance (DESeq2) → ps_deseq: non-rarefied integer counts derived from ps_filt, with optional count-based taxon prefilter
(default: taxa total counts ≥ 10 across all samples)
From ps_filt (e.g. 5669 taxa and 239 samples), we branch into analysis-ready objects in two directions:
Normalize number of reads in each sample using median sequencing depth.
# RAREFACTION
set.seed(9242) # This will help in reproducing the filtering and nomalisation.
ps_rarefied <- rarefy_even_depth(ps_filt, sample.size = 6389)
#total <- 6389
# # NORMALIZE number of reads in each sample using median sequencing depth.
# total = median(sample_sums(ps.ng.tax))
# #> total
# #[1] 42369
# standf = function(x, t=total) round(t * (x / sum(x)))
# ps.ng.tax = transform_sample_counts(ps.ng.tax, standf)
# ps_rel <- microbiome::transform(ps.ng.tax, "compositional")
#
# saveRDS(ps.ng.tax, "./ps.ng.tax.rds")
For the heatmaps, we focus on the most abundant OTUs by first converting counts to relative abundances within each sample. We then filter to retain only OTUs whose mean relative abundance across all samples exceeds 0.1% (0.001). We are left with 199 OTUs which makes the reading much more easy.
# 1) Convert to relative abundances
ps_rel <- transform_sample_counts(ps_filt, function(x) x / sum(x))
# 2) Get the logical vector of which OTUs to keep (based on relative abundance)
keep_vector <- phyloseq::filter_taxa(
ps_rel,
function(x) mean(x) > 0.001,
prune = FALSE
)
# 3) Use the TRUE/FALSE vector to subset absolute abundance data
ps_abund <- prune_taxa(names(keep_vector)[keep_vector], ps_filt)
# 4) Normalize the final subset to relative abundances per sample
ps_abund_rel <- transform_sample_counts(
ps_abund,
function(x) x / sum(x)
)
library(stringr)
#for id in 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206; do
#for id in 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62; do
# echo "phyloseq::tax_table(ps_abund_rel)[${id},\"Domain\"] <- str_split(phyloseq::tax_table(ps_abund_rel)[${id},\"Domain\"], \"__\")[[1]][2]"
# echo "phyloseq::tax_table(ps_abund_rel)[${id},\"Phylum\"] <- str_split(phyloseq::tax_table(ps_abund_rel)[${id},\"Phylum\"], \"__\")[[1]][2]"
# echo "phyloseq::tax_table(ps_abund_rel)[${id},\"Class\"] <- str_split(phyloseq::tax_table(ps_abund_rel)[${id},\"Class\"], \"__\")[[1]][2]"
# echo "phyloseq::tax_table(ps_abund_rel)[${id},\"Order\"] <- str_split(phyloseq::tax_table(ps_abund_rel)[${id},\"Order\"], \"__\")[[1]][2]"
# echo "phyloseq::tax_table(ps_abund_rel)[${id},\"Family\"] <- str_split(phyloseq::tax_table(ps_abund_rel)[${id},\"Family\"], \"__\")[[1]][2]"
# echo "phyloseq::tax_table(ps_abund_rel)[${id},\"Genus\"] <- str_split(phyloseq::tax_table(ps_abund_rel)[${id},\"Genus\"], \"__\")[[1]][2]"
# echo "phyloseq::tax_table(ps_abund_rel)[${id},\"Species\"] <- str_split(phyloseq::tax_table(ps_abund_rel)[${id},\"Species\"], \"__\")[[1]][2]"
#done
phyloseq::tax_table(ps_abund_rel)[1,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[1,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[1,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[1,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[1,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[1,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[1,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[1,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[1,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[1,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[1,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[1,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[1,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[1,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[2,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[2,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[2,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[2,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[2,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[2,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[2,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[2,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[2,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[2,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[2,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[2,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[2,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[2,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[3,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[3,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[3,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[3,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[3,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[3,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[3,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[3,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[3,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[3,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[3,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[3,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[3,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[3,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[4,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[4,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[4,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[4,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[4,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[4,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[4,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[4,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[4,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[4,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[4,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[4,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[4,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[4,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[5,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[5,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[5,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[5,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[5,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[5,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[5,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[5,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[5,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[5,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[5,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[5,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[5,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[5,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[6,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[6,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[6,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[6,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[6,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[6,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[6,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[6,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[6,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[6,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[6,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[6,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[6,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[6,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[7,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[7,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[7,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[7,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[7,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[7,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[7,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[7,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[7,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[7,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[7,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[7,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[7,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[7,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[8,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[8,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[8,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[8,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[8,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[8,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[8,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[8,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[8,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[8,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[8,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[8,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[8,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[8,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[9,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[9,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[9,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[9,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[9,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[9,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[9,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[9,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[9,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[9,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[9,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[9,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[9,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[9,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[10,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[10,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[10,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[10,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[10,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[10,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[10,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[10,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[10,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[10,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[10,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[10,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[10,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[10,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[11,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[11,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[11,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[11,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[11,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[11,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[11,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[11,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[11,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[11,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[11,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[11,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[11,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[11,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[12,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[12,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[12,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[12,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[12,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[12,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[12,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[12,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[12,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[12,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[12,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[12,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[12,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[12,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[13,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[13,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[13,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[13,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[13,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[13,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[13,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[13,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[13,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[13,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[13,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[13,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[13,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[13,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[14,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[14,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[14,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[14,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[14,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[14,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[14,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[14,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[14,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[14,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[14,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[14,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[14,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[14,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[15,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[15,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[15,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[15,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[15,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[15,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[15,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[15,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[15,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[15,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[15,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[15,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[15,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[15,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[16,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[16,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[16,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[16,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[16,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[16,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[16,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[16,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[16,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[16,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[16,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[16,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[16,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[16,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[17,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[17,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[17,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[17,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[17,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[17,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[17,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[17,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[17,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[17,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[17,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[17,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[17,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[17,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[18,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[18,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[18,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[18,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[18,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[18,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[18,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[18,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[18,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[18,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[18,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[18,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[18,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[18,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[19,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[19,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[19,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[19,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[19,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[19,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[19,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[19,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[19,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[19,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[19,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[19,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[19,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[19,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[20,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[20,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[20,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[20,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[20,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[20,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[20,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[20,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[20,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[20,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[20,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[20,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[20,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[20,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[21,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[21,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[21,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[21,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[21,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[21,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[21,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[21,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[21,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[21,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[21,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[21,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[21,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[21,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[22,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[22,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[22,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[22,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[22,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[22,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[22,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[22,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[22,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[22,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[22,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[22,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[22,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[22,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[23,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[23,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[23,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[23,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[23,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[23,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[23,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[23,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[23,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[23,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[23,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[23,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[23,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[23,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[24,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[24,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[24,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[24,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[24,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[24,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[24,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[24,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[24,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[24,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[24,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[24,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[24,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[24,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[25,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[25,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[25,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[25,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[25,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[25,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[25,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[25,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[25,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[25,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[25,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[25,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[25,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[25,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[26,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[26,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[26,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[26,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[26,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[26,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[26,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[26,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[26,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[26,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[26,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[26,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[26,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[26,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[27,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[27,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[27,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[27,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[27,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[27,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[27,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[27,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[27,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[27,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[27,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[27,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[27,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[27,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[28,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[28,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[28,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[28,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[28,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[28,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[28,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[28,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[28,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[28,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[28,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[28,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[28,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[28,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[29,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[29,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[29,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[29,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[29,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[29,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[29,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[29,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[29,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[29,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[29,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[29,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[29,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[29,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[30,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[30,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[30,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[30,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[30,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[30,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[30,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[30,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[30,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[30,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[30,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[30,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[30,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[30,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[31,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[31,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[31,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[31,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[31,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[31,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[31,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[31,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[31,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[31,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[31,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[31,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[31,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[31,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[32,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[32,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[32,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[32,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[32,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[32,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[32,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[32,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[32,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[32,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[32,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[32,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[32,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[32,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[33,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[33,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[33,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[33,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[33,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[33,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[33,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[33,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[33,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[33,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[33,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[33,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[33,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[33,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[34,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[34,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[34,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[34,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[34,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[34,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[34,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[34,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[34,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[34,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[34,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[34,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[34,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[34,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[35,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[35,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[35,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[35,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[35,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[35,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[35,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[35,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[35,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[35,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[35,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[35,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[35,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[35,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[36,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[36,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[36,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[36,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[36,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[36,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[36,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[36,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[36,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[36,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[36,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[36,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[36,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[36,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[37,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[37,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[37,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[37,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[37,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[37,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[37,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[37,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[37,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[37,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[37,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[37,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[37,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[37,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[38,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[38,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[38,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[38,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[38,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[38,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[38,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[38,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[38,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[38,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[38,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[38,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[38,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[38,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[39,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[39,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[39,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[39,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[39,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[39,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[39,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[39,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[39,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[39,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[39,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[39,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[39,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[39,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[40,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[40,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[40,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[40,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[40,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[40,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[40,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[40,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[40,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[40,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[40,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[40,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[40,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[40,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[41,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[41,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[41,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[41,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[41,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[41,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[41,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[41,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[41,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[41,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[41,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[41,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[41,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[41,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[42,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[42,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[42,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[42,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[42,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[42,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[42,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[42,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[42,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[42,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[42,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[42,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[42,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[42,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[43,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[43,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[43,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[43,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[43,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[43,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[43,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[43,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[43,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[43,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[43,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[43,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[43,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[43,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[44,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[44,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[44,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[44,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[44,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[44,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[44,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[44,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[44,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[44,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[44,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[44,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[44,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[44,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[45,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[45,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[45,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[45,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[45,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[45,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[45,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[45,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[45,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[45,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[45,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[45,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[45,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[45,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[46,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[46,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[46,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[46,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[46,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[46,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[46,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[46,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[46,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[46,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[46,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[46,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[46,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[46,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[47,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[47,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[47,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[47,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[47,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[47,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[47,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[47,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[47,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[47,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[47,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[47,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[47,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[47,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[48,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[48,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[48,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[48,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[48,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[48,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[48,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[48,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[48,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[48,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[48,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[48,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[48,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[48,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[49,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[49,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[49,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[49,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[49,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[49,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[49,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[49,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[49,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[49,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[49,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[49,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[49,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[49,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[50,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[50,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[50,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[50,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[50,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[50,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[50,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[50,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[50,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[50,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[50,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[50,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[50,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[50,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[51,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[51,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[51,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[51,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[51,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[51,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[51,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[51,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[51,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[51,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[51,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[51,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[51,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[51,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[52,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[52,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[52,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[52,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[52,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[52,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[52,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[52,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[52,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[52,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[52,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[52,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[52,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[52,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[53,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[53,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[53,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[53,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[53,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[53,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[53,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[53,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[53,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[53,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[53,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[53,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[53,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[53,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[54,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[54,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[54,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[54,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[54,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[54,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[54,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[54,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[54,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[54,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[54,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[54,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[54,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[54,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[55,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[55,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[55,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[55,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[55,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[55,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[55,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[55,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[55,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[55,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[55,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[55,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[55,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[55,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[56,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[56,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[56,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[56,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[56,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[56,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[56,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[56,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[56,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[56,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[56,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[56,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[56,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[56,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[57,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[57,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[57,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[57,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[57,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[57,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[57,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[57,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[57,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[57,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[57,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[57,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[57,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[57,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[58,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[58,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[58,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[58,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[58,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[58,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[58,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[58,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[58,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[58,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[58,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[58,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[58,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[58,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[59,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[59,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[59,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[59,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[59,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[59,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[59,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[59,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[59,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[59,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[59,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[59,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[59,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[59,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[60,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[60,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[60,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[60,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[60,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[60,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[60,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[60,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[60,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[60,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[60,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[60,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[60,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[60,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[61,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[61,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[61,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[61,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[61,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[61,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[61,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[61,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[61,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[61,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[61,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[61,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[61,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[61,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[62,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[62,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[62,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[62,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[62,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[62,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[62,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[62,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[62,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[62,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[62,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[62,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[62,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[62,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[63,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[63,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[63,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[63,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[63,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[63,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[63,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[63,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[63,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[63,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[63,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[63,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[63,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[63,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[64,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[64,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[64,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[64,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[64,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[64,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[64,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[64,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[64,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[64,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[64,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[64,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[64,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[64,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[65,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[65,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[65,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[65,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[65,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[65,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[65,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[65,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[65,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[65,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[65,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[65,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[65,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[65,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[66,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[66,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[66,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[66,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[66,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[66,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[66,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[66,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[66,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[66,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[66,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[66,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[66,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[66,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[67,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[67,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[67,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[67,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[67,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[67,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[67,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[67,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[67,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[67,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[67,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[67,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[67,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[67,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[68,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[68,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[68,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[68,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[68,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[68,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[68,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[68,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[68,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[68,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[68,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[68,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[68,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[68,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[69,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[69,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[69,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[69,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[69,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[69,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[69,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[69,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[69,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[69,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[69,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[69,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[69,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[69,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[70,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[70,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[70,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[70,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[70,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[70,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[70,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[70,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[70,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[70,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[70,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[70,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[70,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[70,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[71,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[71,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[71,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[71,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[71,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[71,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[71,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[71,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[71,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[71,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[71,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[71,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[71,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[71,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[72,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[72,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[72,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[72,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[72,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[72,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[72,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[72,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[72,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[72,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[72,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[72,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[72,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[72,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[73,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[73,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[73,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[73,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[73,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[73,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[73,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[73,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[73,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[73,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[73,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[73,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[73,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[73,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[74,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[74,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[74,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[74,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[74,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[74,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[74,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[74,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[74,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[74,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[74,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[74,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[74,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[74,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[75,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[75,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[75,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[75,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[75,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[75,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[75,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[75,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[75,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[75,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[75,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[75,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[75,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[75,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[76,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[76,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[76,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[76,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[76,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[76,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[76,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[76,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[76,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[76,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[76,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[76,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[76,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[76,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[77,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[77,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[77,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[77,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[77,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[77,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[77,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[77,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[77,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[77,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[77,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[77,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[77,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[77,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[78,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[78,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[78,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[78,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[78,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[78,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[78,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[78,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[78,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[78,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[78,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[78,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[78,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[78,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[79,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[79,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[79,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[79,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[79,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[79,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[79,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[79,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[79,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[79,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[79,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[79,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[79,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[79,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[80,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[80,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[80,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[80,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[80,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[80,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[80,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[80,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[80,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[80,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[80,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[80,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[80,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[80,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[81,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[81,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[81,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[81,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[81,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[81,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[81,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[81,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[81,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[81,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[81,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[81,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[81,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[81,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[82,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[82,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[82,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[82,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[82,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[82,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[82,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[82,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[82,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[82,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[82,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[82,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[82,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[82,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[83,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[83,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[83,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[83,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[83,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[83,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[83,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[83,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[83,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[83,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[83,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[83,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[83,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[83,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[84,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[84,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[84,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[84,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[84,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[84,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[84,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[84,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[84,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[84,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[84,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[84,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[84,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[84,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[85,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[85,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[85,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[85,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[85,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[85,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[85,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[85,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[85,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[85,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[85,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[85,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[85,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[85,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[86,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[86,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[86,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[86,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[86,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[86,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[86,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[86,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[86,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[86,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[86,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[86,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[86,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[86,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[87,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[87,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[87,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[87,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[87,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[87,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[87,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[87,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[87,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[87,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[87,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[87,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[87,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[87,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[88,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[88,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[88,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[88,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[88,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[88,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[88,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[88,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[88,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[88,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[88,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[88,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[88,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[88,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[89,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[89,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[89,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[89,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[89,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[89,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[89,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[89,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[89,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[89,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[89,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[89,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[89,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[89,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[90,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[90,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[90,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[90,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[90,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[90,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[90,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[90,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[90,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[90,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[90,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[90,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[90,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[90,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[91,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[91,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[91,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[91,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[91,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[91,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[91,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[91,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[91,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[91,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[91,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[91,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[91,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[91,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[92,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[92,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[92,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[92,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[92,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[92,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[92,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[92,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[92,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[92,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[92,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[92,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[92,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[92,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[93,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[93,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[93,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[93,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[93,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[93,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[93,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[93,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[93,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[93,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[93,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[93,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[93,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[93,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[94,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[94,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[94,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[94,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[94,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[94,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[94,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[94,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[94,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[94,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[94,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[94,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[94,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[94,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[95,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[95,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[95,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[95,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[95,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[95,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[95,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[95,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[95,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[95,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[95,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[95,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[95,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[95,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[96,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[96,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[96,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[96,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[96,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[96,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[96,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[96,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[96,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[96,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[96,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[96,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[96,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[96,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[97,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[97,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[97,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[97,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[97,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[97,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[97,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[97,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[97,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[97,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[97,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[97,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[97,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[97,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[98,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[98,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[98,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[98,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[98,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[98,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[98,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[98,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[98,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[98,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[98,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[98,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[98,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[98,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[99,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[99,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[99,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[99,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[99,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[99,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[99,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[99,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[99,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[99,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[99,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[99,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[99,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[99,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[100,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[100,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[100,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[100,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[100,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[100,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[100,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[100,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[100,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[100,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[100,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[100,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[100,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[100,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[101,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[101,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[101,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[101,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[101,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[101,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[101,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[101,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[101,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[101,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[101,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[101,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[101,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[101,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[102,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[102,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[102,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[102,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[102,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[102,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[102,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[102,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[102,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[102,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[102,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[102,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[102,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[102,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[103,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[103,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[103,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[103,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[103,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[103,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[103,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[103,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[103,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[103,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[103,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[103,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[103,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[103,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[104,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[104,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[104,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[104,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[104,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[104,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[104,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[104,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[104,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[104,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[104,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[104,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[104,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[104,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[105,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[105,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[105,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[105,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[105,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[105,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[105,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[105,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[105,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[105,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[105,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[105,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[105,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[105,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[106,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[106,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[106,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[106,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[106,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[106,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[106,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[106,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[106,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[106,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[106,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[106,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[106,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[106,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[107,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[107,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[107,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[107,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[107,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[107,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[107,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[107,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[107,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[107,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[107,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[107,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[107,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[107,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[108,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[108,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[108,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[108,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[108,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[108,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[108,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[108,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[108,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[108,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[108,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[108,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[108,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[108,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[109,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[109,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[109,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[109,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[109,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[109,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[109,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[109,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[109,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[109,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[109,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[109,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[109,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[109,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[110,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[110,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[110,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[110,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[110,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[110,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[110,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[110,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[110,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[110,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[110,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[110,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[110,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[110,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[111,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[111,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[111,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[111,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[111,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[111,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[111,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[111,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[111,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[111,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[111,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[111,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[111,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[111,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[112,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[112,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[112,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[112,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[112,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[112,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[112,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[112,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[112,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[112,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[112,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[112,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[112,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[112,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[113,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[113,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[113,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[113,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[113,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[113,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[113,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[113,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[113,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[113,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[113,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[113,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[113,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[113,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[114,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[114,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[114,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[114,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[114,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[114,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[114,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[114,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[114,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[114,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[114,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[114,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[114,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[114,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[115,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[115,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[115,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[115,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[115,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[115,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[115,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[115,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[115,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[115,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[115,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[115,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[115,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[115,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[116,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[116,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[116,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[116,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[116,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[116,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[116,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[116,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[116,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[116,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[116,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[116,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[116,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[116,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[117,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[117,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[117,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[117,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[117,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[117,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[117,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[117,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[117,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[117,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[117,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[117,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[117,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[117,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[118,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[118,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[118,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[118,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[118,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[118,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[118,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[118,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[118,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[118,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[118,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[118,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[118,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[118,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[119,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[119,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[119,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[119,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[119,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[119,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[119,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[119,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[119,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[119,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[119,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[119,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[119,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[119,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[120,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[120,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[120,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[120,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[120,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[120,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[120,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[120,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[120,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[120,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[120,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[120,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[120,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[120,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[121,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[121,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[121,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[121,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[121,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[121,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[121,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[121,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[121,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[121,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[121,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[121,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[121,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[121,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[122,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[122,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[122,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[122,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[122,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[122,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[122,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[122,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[122,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[122,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[122,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[122,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[122,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[122,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[123,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[123,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[123,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[123,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[123,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[123,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[123,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[123,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[123,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[123,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[123,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[123,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[123,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[123,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[124,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[124,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[124,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[124,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[124,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[124,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[124,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[124,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[124,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[124,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[124,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[124,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[124,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[124,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[125,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[125,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[125,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[125,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[125,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[125,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[125,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[125,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[125,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[125,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[125,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[125,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[125,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[125,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[126,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[126,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[126,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[126,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[126,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[126,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[126,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[126,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[126,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[126,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[126,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[126,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[126,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[126,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[127,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[127,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[127,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[127,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[127,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[127,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[127,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[127,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[127,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[127,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[127,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[127,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[127,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[127,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[128,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[128,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[128,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[128,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[128,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[128,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[128,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[128,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[128,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[128,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[128,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[128,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[128,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[128,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[129,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[129,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[129,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[129,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[129,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[129,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[129,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[129,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[129,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[129,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[129,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[129,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[129,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[129,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[130,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[130,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[130,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[130,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[130,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[130,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[130,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[130,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[130,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[130,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[130,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[130,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[130,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[130,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[131,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[131,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[131,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[131,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[131,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[131,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[131,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[131,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[131,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[131,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[131,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[131,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[131,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[131,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[132,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[132,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[132,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[132,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[132,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[132,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[132,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[132,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[132,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[132,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[132,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[132,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[132,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[132,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[133,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[133,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[133,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[133,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[133,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[133,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[133,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[133,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[133,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[133,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[133,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[133,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[133,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[133,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[134,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[134,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[134,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[134,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[134,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[134,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[134,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[134,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[134,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[134,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[134,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[134,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[134,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[134,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[135,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[135,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[135,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[135,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[135,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[135,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[135,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[135,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[135,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[135,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[135,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[135,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[135,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[135,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[136,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[136,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[136,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[136,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[136,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[136,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[136,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[136,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[136,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[136,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[136,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[136,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[136,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[136,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[137,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[137,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[137,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[137,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[137,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[137,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[137,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[137,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[137,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[137,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[137,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[137,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[137,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[137,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[138,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[138,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[138,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[138,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[138,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[138,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[138,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[138,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[138,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[138,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[138,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[138,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[138,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[138,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[139,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[139,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[139,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[139,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[139,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[139,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[139,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[139,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[139,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[139,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[139,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[139,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[139,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[139,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[140,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[140,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[140,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[140,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[140,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[140,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[140,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[140,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[140,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[140,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[140,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[140,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[140,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[140,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[141,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[141,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[141,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[141,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[141,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[141,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[141,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[141,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[141,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[141,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[141,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[141,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[141,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[141,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[142,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[142,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[142,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[142,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[142,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[142,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[142,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[142,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[142,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[142,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[142,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[142,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[142,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[142,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[143,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[143,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[143,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[143,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[143,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[143,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[143,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[143,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[143,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[143,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[143,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[143,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[143,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[143,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[144,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[144,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[144,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[144,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[144,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[144,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[144,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[144,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[144,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[144,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[144,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[144,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[144,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[144,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[145,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[145,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[145,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[145,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[145,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[145,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[145,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[145,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[145,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[145,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[145,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[145,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[145,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[145,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[146,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[146,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[146,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[146,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[146,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[146,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[146,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[146,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[146,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[146,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[146,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[146,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[146,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[146,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[147,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[147,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[147,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[147,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[147,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[147,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[147,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[147,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[147,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[147,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[147,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[147,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[147,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[147,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[148,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[148,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[148,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[148,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[148,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[148,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[148,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[148,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[148,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[148,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[148,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[148,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[148,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[148,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[149,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[149,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[149,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[149,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[149,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[149,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[149,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[149,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[149,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[149,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[149,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[149,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[149,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[149,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[150,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[150,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[150,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[150,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[150,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[150,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[150,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[150,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[150,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[150,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[150,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[150,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[150,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[150,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[151,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[151,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[151,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[151,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[151,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[151,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[151,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[151,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[151,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[151,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[151,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[151,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[151,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[151,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[152,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[152,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[152,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[152,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[152,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[152,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[152,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[152,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[152,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[152,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[152,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[152,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[152,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[152,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[153,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[153,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[153,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[153,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[153,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[153,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[153,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[153,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[153,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[153,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[153,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[153,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[153,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[153,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[154,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[154,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[154,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[154,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[154,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[154,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[154,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[154,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[154,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[154,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[154,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[154,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[154,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[154,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[155,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[155,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[155,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[155,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[155,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[155,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[155,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[155,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[155,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[155,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[155,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[155,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[155,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[155,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[156,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[156,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[156,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[156,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[156,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[156,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[156,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[156,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[156,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[156,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[156,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[156,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[156,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[156,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[157,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[157,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[157,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[157,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[157,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[157,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[157,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[157,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[157,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[157,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[157,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[157,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[157,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[157,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[158,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[158,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[158,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[158,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[158,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[158,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[158,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[158,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[158,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[158,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[158,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[158,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[158,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[158,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[159,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[159,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[159,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[159,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[159,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[159,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[159,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[159,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[159,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[159,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[159,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[159,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[159,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[159,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[160,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[160,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[160,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[160,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[160,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[160,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[160,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[160,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[160,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[160,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[160,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[160,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[160,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[160,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[161,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[161,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[161,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[161,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[161,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[161,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[161,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[161,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[161,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[161,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[161,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[161,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[161,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[161,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[162,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[162,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[162,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[162,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[162,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[162,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[162,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[162,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[162,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[162,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[162,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[162,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[162,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[162,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[163,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[163,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[163,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[163,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[163,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[163,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[163,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[163,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[163,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[163,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[163,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[163,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[163,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[163,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[164,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[164,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[164,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[164,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[164,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[164,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[164,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[164,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[164,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[164,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[164,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[164,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[164,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[164,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[165,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[165,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[165,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[165,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[165,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[165,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[165,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[165,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[165,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[165,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[165,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[165,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[165,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[165,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[166,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[166,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[166,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[166,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[166,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[166,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[166,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[166,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[166,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[166,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[166,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[166,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[166,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[166,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[167,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[167,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[167,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[167,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[167,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[167,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[167,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[167,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[167,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[167,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[167,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[167,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[167,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[167,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[168,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[168,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[168,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[168,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[168,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[168,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[168,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[168,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[168,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[168,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[168,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[168,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[168,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[168,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[169,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[169,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[169,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[169,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[169,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[169,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[169,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[169,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[169,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[169,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[169,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[169,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[169,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[169,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[170,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[170,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[170,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[170,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[170,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[170,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[170,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[170,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[170,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[170,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[170,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[170,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[170,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[170,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[171,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[171,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[171,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[171,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[171,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[171,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[171,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[171,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[171,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[171,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[171,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[171,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[171,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[171,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[172,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[172,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[172,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[172,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[172,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[172,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[172,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[172,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[172,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[172,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[172,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[172,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[172,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[172,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[173,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[173,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[173,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[173,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[173,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[173,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[173,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[173,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[173,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[173,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[173,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[173,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[173,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[173,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[174,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[174,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[174,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[174,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[174,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[174,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[174,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[174,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[174,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[174,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[174,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[174,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[174,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[174,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[175,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[175,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[175,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[175,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[175,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[175,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[175,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[175,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[175,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[175,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[175,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[175,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[175,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[175,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[176,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[176,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[176,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[176,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[176,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[176,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[176,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[176,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[176,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[176,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[176,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[176,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[176,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[176,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[177,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[177,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[177,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[177,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[177,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[177,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[177,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[177,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[177,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[177,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[177,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[177,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[177,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[177,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[178,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[178,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[178,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[178,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[178,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[178,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[178,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[178,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[178,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[178,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[178,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[178,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[178,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[178,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[179,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[179,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[179,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[179,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[179,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[179,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[179,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[179,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[179,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[179,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[179,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[179,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[179,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[179,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[180,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[180,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[180,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[180,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[180,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[180,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[180,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[180,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[180,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[180,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[180,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[180,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[180,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[180,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[181,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[181,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[181,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[181,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[181,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[181,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[181,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[181,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[181,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[181,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[181,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[181,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[181,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[181,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[182,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[182,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[182,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[182,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[182,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[182,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[182,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[182,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[182,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[182,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[182,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[182,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[182,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[182,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[183,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[183,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[183,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[183,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[183,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[183,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[183,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[183,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[183,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[183,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[183,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[183,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[183,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[183,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[184,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[184,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[184,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[184,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[184,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[184,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[184,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[184,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[184,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[184,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[184,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[184,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[184,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[184,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[185,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[185,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[185,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[185,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[185,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[185,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[185,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[185,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[185,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[185,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[185,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[185,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[185,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[185,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[186,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[186,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[186,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[186,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[186,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[186,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[186,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[186,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[186,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[186,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[186,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[186,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[186,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[186,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[187,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[187,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[187,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[187,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[187,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[187,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[187,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[187,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[187,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[187,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[187,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[187,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[187,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[187,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[188,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[188,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[188,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[188,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[188,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[188,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[188,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[188,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[188,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[188,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[188,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[188,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[188,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[188,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[189,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[189,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[189,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[189,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[189,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[189,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[189,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[189,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[189,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[189,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[189,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[189,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[189,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[189,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[190,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[190,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[190,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[190,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[190,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[190,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[190,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[190,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[190,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[190,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[190,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[190,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[190,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[190,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[191,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[191,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[191,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[191,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[191,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[191,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[191,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[191,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[191,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[191,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[191,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[191,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[191,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[191,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[192,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[192,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[192,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[192,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[192,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[192,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[192,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[192,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[192,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[192,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[192,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[192,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[192,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[192,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[193,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[193,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[193,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[193,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[193,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[193,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[193,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[193,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[193,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[193,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[193,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[193,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[193,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[193,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[194,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[194,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[194,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[194,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[194,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[194,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[194,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[194,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[194,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[194,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[194,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[194,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[194,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[194,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[195,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[195,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[195,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[195,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[195,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[195,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[195,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[195,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[195,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[195,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[195,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[195,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[195,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[195,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[196,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[196,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[196,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[196,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[196,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[196,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[196,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[196,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[196,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[196,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[196,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[196,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[196,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[196,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[197,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[197,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[197,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[197,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[197,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[197,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[197,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[197,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[197,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[197,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[197,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[197,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[197,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[197,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[198,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[198,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[198,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[198,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[198,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[198,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[198,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[198,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[198,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[198,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[198,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[198,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[198,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[198,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[199,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[199,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[199,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[199,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[199,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[199,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[199,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[199,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[199,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[199,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[199,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[199,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[199,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[199,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[200,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[200,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[200,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[200,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[200,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[200,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[200,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[200,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[200,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[200,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[200,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[200,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[200,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[200,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[201,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[201,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[201,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[201,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[201,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[201,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[201,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[201,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[201,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[201,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[201,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[201,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[201,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[201,"Species"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[202,"Domain"] <- str_split(phyloseq::tax_table(ps_abund_rel)[202,"Domain"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[202,"Phylum"] <- str_split(phyloseq::tax_table(ps_abund_rel)[202,"Phylum"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[202,"Class"] <- str_split(phyloseq::tax_table(ps_abund_rel)[202,"Class"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[202,"Order"] <- str_split(phyloseq::tax_table(ps_abund_rel)[202,"Order"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[202,"Family"] <- str_split(phyloseq::tax_table(ps_abund_rel)[202,"Family"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[202,"Genus"] <- str_split(phyloseq::tax_table(ps_abund_rel)[202,"Genus"], "__")[[1]][2]
phyloseq::tax_table(ps_abund_rel)[202,"Species"] <- str_split(phyloseq::tax_table(ps_abund_rel)[202,"Species"], "__")[[1]][2]
Taxonomic summary
Bar plots in phylum level
#aes(color="Phylum", fill="Phylum") --> aes()
#ggplot(data=data, aes(x=Sample, y=Abundance, fill=Phylum))
#options(max.print = 1e6)
my_colors <- c("darkblue", "darkgoldenrod1", "darkseagreen", "darkorchid", "darkolivegreen1", "lightskyblue", "darkgreen", "deeppink", "khaki2", "firebrick", "brown1", "darkorange1", "cyan1", "royalblue4", "darksalmon", "darkblue","royalblue4", "dodgerblue3", "steelblue1", "lightskyblue", "darkseagreen", "darkgoldenrod1", "darkseagreen", "darkorchid", "darkolivegreen1", "brown1", "darkorange1", "cyan1", "darkgrey")
plot_bar(ps_abund_rel, fill="Phylum") + geom_bar(aes(), stat="identity", position="stack") +
scale_fill_manual(values = my_colors) + theme(axis.text = element_text(size = 7, colour="black")) + theme(legend.position="bottom") + guides(fill=guide_legend(nrow=2)) #6 instead of theme.size
\pagebreak
Regroup together pre vs post stroke samples and normalize number of reads in each group using median sequencing depth.
ps_abund_rel_group <- merge_samples(ps_abund_rel, "G9_10_11_preFMT")
#PENDING: The effect weighted twice by sum(x), is the same to the effect weighted once directly from absolute abundance?!
ps_abund_rel_group_ = transform_sample_counts(ps_abund_rel_group, function(x) x / sum(x))
#plot_bar(ps_abund_relSampleType_, fill = "Phylum") + geom_bar(aes(color=Phylum, fill=Phylum), stat="identity", position="stack")
plot_bar(ps_abund_rel_group_, fill="Phylum") +
geom_bar(stat="identity", position="stack") +
scale_x_discrete(limits = c("pre-FMT", "Group9","Group10","Group11")) +
scale_fill_manual(values = my_colors) +
theme(
axis.text.x = element_text(angle = 30, hjust = 1, vjust = 1, size = 11, colour = "black"),
axis.text.y = element_text(angle = 0, size = 11, colour = "black"),
axis.title.x = element_text(size = 12, colour = "black"),
axis.title.y = element_text(size = 12, colour = "black")
)
Bar plots in class level
my_colors <- c("darkblue", "darkgoldenrod1", "darkseagreen", "darkorchid", "darkolivegreen1", "lightskyblue", "darkgreen", "deeppink", "khaki2", "firebrick", "brown1", "darkorange1", "cyan1", "royalblue4", "darksalmon", "darkblue","royalblue4", "dodgerblue3", "steelblue1", "lightskyblue", "darkseagreen", "darkgoldenrod1", "darkseagreen", "darkorchid", "darkolivegreen1", "brown1", "darkorange1", "cyan1", "darkgrey")
plot_bar(ps_abund_rel, fill="Class") + geom_bar(aes(), stat="identity", position="stack") +
scale_fill_manual(values = my_colors) + theme(axis.text = element_text(size = 7, colour="black")) + theme(legend.position="bottom") + guides(fill=guide_legend(nrow=3))
Regroup together pre vs post stroke samples and normalize number of reads in each group using median sequencing depth.
plot_bar(ps_abund_rel_group_, fill="Class") +
geom_bar(stat="identity", position="stack") +
scale_x_discrete(limits = c("pre-FMT", "Group9","Group10","Group11")) +
scale_fill_manual(values = my_colors) +
theme(
axis.text.x = element_text(angle = 30, hjust = 1, vjust = 1, size = 11, colour = "black"),
axis.text.y = element_text(angle = 0, size = 11, colour = "black"),
axis.title.x = element_text(size = 12, colour = "black"),
axis.title.y = element_text(size = 12, colour = "black")
)
\pagebreak
Bar plots in order level
my_colors <- c("darkblue", "darkgoldenrod1", "darkseagreen", "darkorchid", "darkolivegreen1", "lightskyblue", "darkgreen", "deeppink", "khaki2", "firebrick", "brown1", "darkorange1", "cyan1", "royalblue4", "darksalmon", "darkblue","royalblue4", "dodgerblue3", "steelblue1", "lightskyblue", "darkseagreen", "darkgoldenrod1", "darkseagreen", "darkorchid", "darkolivegreen1", "brown1", "darkorange1", "cyan1", "darkgrey")
plot_bar(ps_abund_rel, fill="Order") + geom_bar(aes(), stat="identity", position="stack") +
scale_fill_manual(values = my_colors) + theme(axis.text = element_text(size = 7, colour="black")) + theme(legend.position="bottom") + guides(fill=guide_legend(nrow=4))
Regroup together pre vs post stroke and normalize number of reads in each group using median sequencing depth.
plot_bar(ps_abund_rel_group_, fill="Order") +
geom_bar(stat="identity", position="stack") +
scale_x_discrete(limits = c("pre-FMT", "Group9","Group10","Group11")) +
scale_fill_manual(values = my_colors) +
theme(
axis.text.x = element_text(angle = 30, hjust = 1, vjust = 1, size = 11, colour = "black"),
axis.text.y = element_text(angle = 0, size = 11, colour = "black"),
axis.title.x = element_text(size = 12, colour = "black"),
axis.title.y = element_text(size = 12, colour = "black")
)
\pagebreak
Bar plots in family level
my_colors <- c(
"#FF0000", "#000000", "#0000FF", "#C0C0C0", "#FFFFFF", "#FFFF00", "#00FFFF", "#FFA500", "#00FF00", "#808080", "#FF00FF", "#800080", "#FDD017", "#0000A0", "#3BB9FF", "#008000", "#800000", "#ADD8E6", "#F778A1", "#800517", "#736F6E", "#F52887", "#C11B17", "#5CB3FF", "#A52A2A", "#FF8040", "#2B60DE", "#736AFF", "#1589FF", "#98AFC7", "#8D38C9", "#307D7E", "#F6358A", "#151B54", "#6D7B8D", "#FDEEF4", "#FF0080", "#F88017", "#2554C7", "#FFF8C6", "#D4A017", "#306EFF", "#151B8D", "#9E7BFF", "#EAC117", "#E0FFFF", "#15317E", "#6C2DC7", "#FBB917", "#FCDFFF", "#15317E", "#254117", "#FAAFBE", "#357EC7"
)
plot_bar(ps_abund_rel, fill="Family") + geom_bar(aes(), stat="identity", position="stack") +
scale_fill_manual(values = my_colors) + theme(axis.text = element_text(size = 7, colour="black")) + theme(legend.position="bottom") + guides(fill=guide_legend(nrow=8))
Regroup together pre vs post stroke samples and normalize number of reads in each group using median sequencing depth.
plot_bar(ps_abund_rel_group_, fill="Family") +
geom_bar(stat="identity", position="stack") +
scale_x_discrete(limits = c("pre-FMT", "Group9","Group10","Group11")) +
scale_fill_manual(values = my_colors) +
theme(
axis.text.x = element_text(angle = 30, hjust = 1, vjust = 1, size = 11, colour = "black"),
axis.text.y = element_text(angle = 0, size = 11, colour = "black"),
axis.title.x = element_text(size = 12, colour = "black"),
axis.title.y = element_text(size = 12, colour = "black")
)
\pagebreak
\pagebreak
Alpha diversity
Plot Chao1 richness estimator, Observed OTUs, Shannon index, and Phylogenetic diversity.
Regroup together samples from the same group.
# using rarefied data
#gunzip table_even4753.biom.gz
#alpha_diversity.py -i table_even42369.biom --metrics chao1,observed_otus,shannon,PD_whole_tree -o adiv_even.txt -t ../clustering/rep_set.tre
#gunzip table_even4753.biom.gz
#alpha_diversity.py -i table_even4753.biom --metrics chao1,observed_otus,shannon,PD_whole_tree -o adiv_even.txt -t ../clustering_stool/rep_set.tre
#gunzip table_even4753.biom.gz
#alpha_diversity.py -i table_even4753.biom --metrics chao1,observed_otus,shannon,PD_whole_tree -o adiv_even.txt -t ../clustering_swab/rep_set.tre
library(dplyr)
library(reshape2)
library(ggpubr)
library(phyloseq)
library(kableExtra)
# ------------------------------------------------------------
# 0) Restrict analysis to the specified sample subset
# ------------------------------------------------------------
stopifnot(exists("samples"))
stopifnot(exists("ps_rarefied"))
ps_rarefied_sub <- prune_samples(samples, ps_rarefied)
hmp.meta <- meta(ps_rarefied_sub)
hmp.meta$sam_name <- rownames(hmp.meta)
# ------------------------------------------------------------
# 1) Read QIIME2 alpha-diversity outputs
# ------------------------------------------------------------
shannon <- read.table("exported_alpha/shannon/alpha-diversity.tsv",
header = TRUE, sep = "\t", stringsAsFactors = FALSE)
faith_pd <- read.table("exported_alpha/faith_pd/alpha-diversity.tsv",
header = TRUE, sep = "\t", stringsAsFactors = FALSE)
observed <- read.table("exported_alpha/observed_features/alpha-diversity.tsv",
header = TRUE, sep = "\t", stringsAsFactors = FALSE)
colnames(shannon) <- c("sam_name", "shannon")
colnames(faith_pd) <- c("sam_name", "PD_whole_tree")
colnames(observed) <- c("sam_name", "observed_otus")
div.df <- Reduce(function(x, y) merge(x, y, by = "sam_name", all = FALSE),
list(shannon, faith_pd, observed))
# ------------------------------------------------------------
# 2) Merge in metadata (from the pruned phyloseq object)
# ------------------------------------------------------------
div.df <- merge(div.df, hmp.meta, by = "sam_name", all.x = TRUE, all.y = FALSE)
# Keep ONLY samples from the predefined list (safety net)
div.df <- div.df %>% filter(sam_name %in% samples)
# ------------------------------------------------------------
# 3) Collapse groups: Group6/7/8 -> pre-FMT; keep only 4 groups
# ------------------------------------------------------------
div.df <- div.df %>%
mutate(
Group4 = case_when(
Group %in% c("Group6","Group7","Group8") ~ "pre-FMT",
Group %in% c("Group9","Group10","Group11") ~ as.character(Group),
TRUE ~ NA_character_
),
Group4 = factor(Group4, levels = c("pre-FMT","Group9","Group10","Group11"))
) %>%
filter(!is.na(Group4))
# ------------------------------------------------------------
# 4) Reformat table for reporting
# ------------------------------------------------------------
div.df2 <- div.df[, c("sam_name", "Group4", "shannon", "observed_otus", "PD_whole_tree")]
colnames(div.df2) <- c("Sample name", "Group", "Shannon", "OTU", "Phylogenetic Diversity")
write.csv(div.df2, file = "alpha_diversities_G9_10_11_preFMT.csv", row.names = FALSE)
#knitr::kable(div.df2) %>%
# kable_styling(bootstrap_options = c("striped","hover","condensed","responsive"))
# ------------------------------------------------------------
# 5) QC: print which samples were used
# ------------------------------------------------------------
requested_samples <- samples
present_in_ps <- sample_names(ps_rarefied)
missing_in_ps <- setdiff(requested_samples, present_in_ps)
cat("\n==== QC: Sample selection (ps_rarefied) ====\n")
cat("Requested samples:", length(requested_samples), "\n")
cat("Present in ps_rarefied:", length(present_in_ps), "\n")
cat("Missing from ps_rarefied:", length(missing_in_ps), "\n")
if (length(missing_in_ps) > 0) print(missing_in_ps)
used_samples <- div.df2$`Sample name`
missing_in_alpha <- setdiff(requested_samples, used_samples)
extra_in_alpha <- setdiff(used_samples, requested_samples)
cat("\n==== QC: Samples used in alpha-div df (div.df2) ====\n")
cat("Used samples:", length(used_samples), "\n")
cat("Requested but NOT used:", length(missing_in_alpha), "\n")
if (length(missing_in_alpha) > 0) print(missing_in_alpha)
cat("Used but NOT requested (should be 0):", length(extra_in_alpha), "\n")
if (length(extra_in_alpha) > 0) print(extra_in_alpha)
qc_table <- div.df %>%
select(sam_name, Group, Group4) %>%
distinct() %>%
arrange(Group4, Group, sam_name)
#cat("\n==== QC: Sample -> Group mapping used for plotting ====\n")
#print(qc_table)
#
#cat("\n==== QC: Counts per collapsed group (Group4) ====\n")
#print(div.df$Group4)
# ------------------------------------------------------------
# 6) Melt + plot
# ------------------------------------------------------------
div_df_melt <- reshape2::melt(
div.df2,
id.vars = c("Sample name","Group"),
variable.name = "variable",
value.name = "value"
)
# ensure final group order
div_df_melt$Group <- factor(div_df_melt$Group, levels = c("pre-FMT","Group9","Group10","Group11"))
#p <- ggboxplot(div_df_melt, x = "Group", y = "value",
# facet.by = "variable",
# scales = "free",
# width = 0.5,
# fill = "gray", legend = "right") +
# theme(
# axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1, size = 8),
# axis.text.y = element_text(size = 10),
# axis.title.x = element_text(size = 12),
# axis.title.y = element_text(size = 12)
# )
# all pairwise comparisons among the 4 groups
#lev <- levels(droplevels(div_df_melt$Group))
#L.pairs <- combn(lev, 2, simplify = FALSE)
#
#p2 <- p + stat_compare_means(
# method = "wilcox.test",
# comparisons = L.pairs,
# label = "p.signif",
# p.adjust.method = "BH",
# hide.ns = FALSE,
# symnum.args = list(
# cutpoints = c(0, 0.0001, 0.001, 0.01, 0.05, 1),
# symbols = c("****", "***", "**", "*", "ns")
# )
#)
#p2
# ------------------------------------------------------------
# 7) Save figure
# ------------------------------------------------------------
#ggsave("./figures/alpha_diversity_G9_10_11_preFMT.png", p2, device="png", height = 10, width = 15, dpi = 300)
#ggsave("./figures/alpha_diversity_G9_10_11_preFMT.svg", p2, device="svg", height = 10, width = 15)
# ------------------------------------------------------------
# 8) Save statistics
# ------------------------------------------------------------
library(dplyr)
library(purrr)
# same pair list as in the plot
lev <- levels(droplevels(div_df_melt$Group))
L.pairs <- combn(lev, 2, simplify = FALSE)
# Pairwise Wilcoxon per metric, with BH adjustment (matches p.adjust.method="BH")
stats_for_plot <- div_df_melt %>%
group_by(variable) %>%
group_modify(~{
dat <- .x
res <- purrr::map_dfr(L.pairs, function(pr) {
g1 <- pr[1]; g2 <- pr[2]
x1 <- dat$value[dat$Group == g1]
x2 <- dat$value[dat$Group == g2]
wt <- suppressWarnings(wilcox.test(x1, x2, exact = FALSE))
tibble(
group1 = g1,
group2 = g2,
n1 = length(x1),
n2 = length(x2),
p = unname(wt$p.value)
)
})
res %>%
mutate(
p.adj = p.adjust(p, method = "BH"),
p.signif = case_when(
p.adj <= 0.0001 ~ "****",
p.adj <= 0.001 ~ "***",
p.adj <= 0.01 ~ "**",
p.adj <= 0.05 ~ "*",
TRUE ~ "ns"
)
)
}) %>%
ungroup() %>%
rename(Metric = variable) %>%
arrange(Metric, group1, group2)
# Print the exact statistics used for annotation
print(stats_for_plot, n = Inf)
write.csv(stats_for_plot, "./figures/alpha_diversity_pairwise_stats_used_for_plot.csv", row.names = FALSE)
library(openxlsx)
# (optional) make sure the folder exists
dir.create("./figures", showWarnings = FALSE, recursive = TRUE)
out_xlsx <- "./figures/alpha_diversity_pairwise_stats_used_for_plot.xlsx"
write.xlsx(
x = list("pairwise_stats" = stats_for_plot),
file = out_xlsx,
overwrite = TRUE
)
cat("Saved Excel file to: ", out_xlsx, "\n", sep = "")
# ---------------------------------------------------------------------------------
# ----------------------------- draw plots from input -----------------------------
library(dplyr)
library(ggpubr)
library(reshape2)
# Example: read from a CSV file you wrote earlier
# stats_for_plot <- read.csv("./figures/alpha_diversity_pairwise_stats_used_for_plot.csv", stringsAsFactors = FALSE)
# Or if you paste as text:
txt <- 'Metric,group1,group2,n1,n2,p,p.adj,p.signif
Shannon,Group10,Group11,6,5,1,1,ns
Shannon,Group9,Group10,6,6,0.297953061608168,0.595906123216336,ns
Shannon,Group9,Group11,6,5,0.522816653919089,0.631819188094748,ns
Shannon,pre-FMT,Group10,18,6,0.017949090591566,0.0538472717746981,ns
Shannon,pre-FMT,Group11,18,5,0.00651735384842052,0.0391041230905231,*
Shannon,pre-FMT,Group9,18,6,0.526515990078957,0.631819188094748,ns
OTU,Group10,Group11,6,5,0.143214612017615,0.214821918026422,ns
OTU,Group9,Group10,6,6,0.229766270461138,0.275719524553366,ns
OTU,Group9,Group11,6,5,0.522816653919089,0.522816653919089,ns
OTU,pre-FMT,Group10,18,6,0.0015274145728676,0.00916448743720559,**
OTU,pre-FMT,Group11,18,5,0.0675601586409843,0.135120317281969,ns
OTU,pre-FMT,Group9,18,6,0.00759484512604444,0.0227845353781333,*
Phylogenetic Diversity,Group10,Group11,6,5,1,1,ns
Phylogenetic Diversity,Group9,Group10,6,6,1,1,ns
Phylogenetic Diversity,Group9,Group11,6,5,0.927264473525232,1,ns
Phylogenetic Diversity,pre-FMT,Group10,18,6,0.000361550896811121,0.00108465269043336,**
Phylogenetic Diversity,pre-FMT,Group11,18,5,0.000910436385913111,0.00182087277182622,**
Phylogenetic Diversity,pre-FMT,Group9,18,6,0.000361550896811121,0.00108465269043336,**'
stats_for_plot <- read.csv(textConnection(txt), stringsAsFactors = FALSE)
# Match naming to your plot data: your facet variable is called "variable"
stats_for_plot <- stats_for_plot %>%
rename(variable = Metric) %>%
mutate(
group1 = factor(group1, levels = c("pre-FMT","Group9","Group10","Group11")),
group2 = factor(group2, levels = c("pre-FMT","Group9","Group10","Group11"))
)
# ------------------------------------------------------------
# Melt + plot (same as yours)
# ------------------------------------------------------------
div_df_melt <- reshape2::melt(
div.df2,
id.vars = c("Sample name","Group"),
variable.name = "variable",
value.name = "value"
)
div_df_melt$Group <- factor(div_df_melt$Group, levels = c("pre-FMT","Group9","Group10","Group11"))
p <- ggboxplot(div_df_melt, x = "Group", y = "value",
facet.by = "variable",
scales = "free",
width = 0.5,
fill = "gray", legend = "right") +
theme(
axis.text.x = element_text(angle = 45, hjust = 1, vjust = 1, size = 8),
axis.text.y = element_text(size = 10),
axis.title.x = element_text(size = 12),
axis.title.y = element_text(size = 12)
)
# ------------------------------------------------------------
# Add significance from your table (NO recomputation)
# ------------------------------------------------------------
# Compute y positions per facet so brackets don't overlap
ymax_by_metric <- div_df_melt %>%
group_by(variable) %>%
summarise(ymax = max(value, na.rm = TRUE), .groups = "drop")
stats_for_plot2 <- stats_for_plot %>%
left_join(ymax_by_metric, by = "variable") %>%
group_by(variable) %>%
arrange(p.adj) %>%
mutate(
# stack brackets within each facet
y.position = ymax * (1.05 + 0.08 * (row_number() - 1)),
# what to display (stars or "ns")
label = p.signif
) %>%
ungroup()
p2 <- p +
stat_pvalue_manual(
stats_for_plot2,
label = "label",
xmin = "group1",
xmax = "group2",
y.position = "y.position",
tip.length = 0.01,
hide.ns = FALSE
)
p2
# ------------------------------------------------------------
# Save
# ------------------------------------------------------------
ggsave("./figures/alpha_diversity_G9_10_11_preFMT_from_table.png",
p2, device="png", height = 6, width = 9, dpi = 300)
ggsave("./figures/alpha_diversity_G9_10_11_preFMT_from_table.svg",
p2, device="svg", height = 6, width = 9)
# label = sprintf("BH p=%.3g", p.adj)
Beta-diversity
Pairwise PERMANOVA tests
Pairwise PERMANOVA tests were performed on Bray–Curtis distance matrices to compare bacterial community composition between all pairs of sample groups (metadata column Group). For each pairwise comparison, the distance matrix was subset to samples from the two groups only, and significance was assessed using vegan::adonis2 with 9,999 permutations. Resulting p-values were adjusted for multiple testing using both Benjamini–Hochberg (BH/FDR) and Bonferroni corrections.
Bray_pairwise_PERMANOVA <- read.csv("figures_MP_Group9_10_11_PreFMT/Bray_pairwise_PERMANOVA.csv", sep = ",")
knitr::kable(Bray_pairwise_PERMANOVA, caption = "Pairwise PERMANOVA results (distance-based community differences among Group levels).") %>%
kable_styling(bootstrap_options = c("striped", "hover", "condensed", "responsive"))
Global PERMANOVA on the weighted UniFrac distance matrix
Groups 9, 10, 11 vs the mixed pre-FMT group
# --- Global beta-diversity (PERMANOVA) ---
cat("```text\n")
cat(
"[PERMANOVA result]\n",
"The object contained internal attribute: PCoA ADONIS\n",
"Permutation test for adonis under reduced model\n",
"Permutation: free\n",
"Number of permutations: 9999\n",
"\n",
"vegan::adonis2(formula = .formula, data = sampleda, permutations = permutations, method = distmethod)\n",
" Df SumOfSqs R2 F Pr(>F) \n",
"Model 3 5.3256 0.58559 14.602 1e-04 ***\n",
"Residual 31 3.7689 0.41441 \n",
"Total 34 9.0945 1.00000 \n",
"---\n",
"Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1\n",
sep = ""
)
cat("```\n")
Groups 9, 10, 11
# --- Global beta-diversity (PERMANOVA) ---
cat("```text\n")
cat(
"[PERMANOVA result]\n",
"The object contained internal attribute: PCoA ADONIS\n",
"Permutation test for adonis under reduced model\n",
"Permutation: free\n",
"Number of permutations: 9999\n",
"\n",
"vegan::adonis2(formula = .formula, data = sampleda, permutations = permutations, method = distmethod)\n",
" Df SumOfSqs R2 F Pr(>F) \n",
"Model 2 0.82208 0.3324 3.4853 1e-04 ***\n",
"Residual 14 1.65109 0.6676 \n",
"Total 16 2.47317 1.0000 \n",
"---\n",
"Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1\n",
sep = ""
)
cat("```\n")
Principal coordinates analysis (PCoA) based on Bray–Curtis dissimilarity
Groups 9, 10, 11 vs the mixed pre-FMT group
knitr::include_graphics("figures_MP_Group9_10_11_PreFMT/PCoA3.png")
Groups 9, 10, 11
knitr::include_graphics("figures_MP_Group9_10_11/PCoA3.png")
Differential abundance analysis
Differential abundance analysis aims to find the differences in the abundance of each taxa between two groups of samples, assigning a significance value to each comparison.
Group 9 vs Group 10
# ------------------------------------------------------------
# DESeq2: non-rarefied integer counts + optional taxon prefilter
# ------------------------------------------------------------
ps_deseq <- ps_filt
ps_deseq_sel2 <- data.table::copy(ps_deseq)
otu_table(ps_deseq_sel2) <- otu_table(ps_deseq)[,c("sample-J1","sample-J2","sample-J3","sample-J4","sample-J10","sample-J11", "sample-K1","sample-K2","sample-K3","sample-K4","sample-K5","sample-K6")]
diagdds = phyloseq_to_deseq2(ps_deseq_sel2, ~Group)
diagdds$Group <- relevel(diagdds$Group, "Group9")
diagdds = DESeq(diagdds, test="Wald", fitType="parametric")
resultsNames(diagdds)
res = results(diagdds, cooksCutoff = FALSE)
alpha = 0.05
sigtab = res[which(res$padj < alpha), ]
sigtab = cbind(as(sigtab, "data.frame"), as(phyloseq::tax_table(ps_deseq_sel2)[rownames(sigtab), ], "matrix"))
# file base name
fname <- "DEGs_Group9_vs_Group10"
write.xlsx(sigtab, file = paste0(fname, ".xlsx"), rowNames = TRUE)
kable(sigtab) %>%
kable_styling(bootstrap_options = c("striped", "hover", "condensed", "responsive"))
library("ggplot2")
theme_set(theme_bw())
scale_fill_discrete <- function(palname = "Set1", ...) {
scale_fill_brewer(palette = palname, ...)
}
x = tapply(sigtab$log2FoldChange, sigtab$Order, function(x) max(x))
x = sort(x)
sigtab$Order = factor(as.character(sigtab$Order), levels=names(x))
x = tapply(sigtab$log2FoldChange, sigtab$Family, function(x) max(x))
x = sort(x)
sigtab$Family = factor(as.character(sigtab$Family), levels=names(x))
#ggplot(sigtab, aes(x=log2FoldChange, y=Family, color=Order)) + geom_point(aes(size=padj)) + scale_size_continuous(name="padj",range=c(8,4))+
theme(axis.text.x = element_text(angle = -25, hjust = 0, vjust=0.5))
# build the plot
p <- ggplot(sigtab, aes(x = log2FoldChange, y = Family, color = Order)) +
geom_point(aes(size = padj)) +
scale_size_continuous(name = "padj", range = c(8, 4)) +
theme_bw() +
theme(axis.text.x = element_text(angle = -25, hjust = 0, vjust = 0.5))
# SVG (svglite gives crisp text)
if (!requireNamespace("svglite", quietly = TRUE)) install.packages("svglite")
ggplot2::ggsave(paste0(fname, ".svg"), plot = p, device = svglite::svglite,
width = 8, height = 6, units = "in", dpi = 300)
# PNG
ggplot2::ggsave(paste0(fname, ".png"), plot = p, device = "png",
width = 8, height = 6, units = "in", dpi = 300)
knitr::include_graphics("DEGs_Group9_vs_Group10.png")