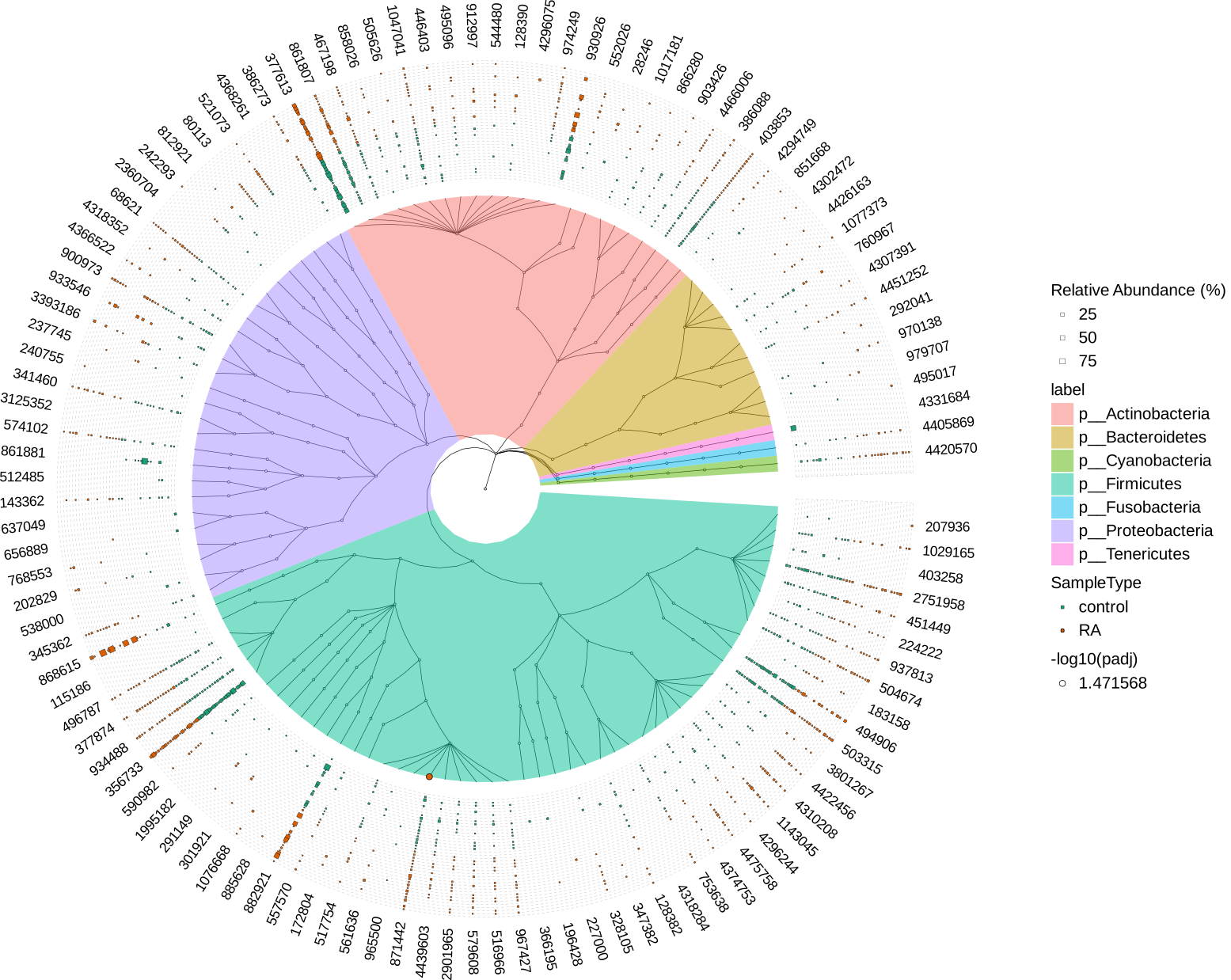
MicrobiotaProcess for RA vs control
gene_x 0 like s 698 view s
Tags:
https://bioconductor.org/packages/release/bioc/vignettes/MicrobiotaProcess/inst/doc//MicrobiotaProcess.html
1, prepare the R environment
#Rscript MicrobiotaProcess.R
#NOTE: exit R script, then login again R-environment; rm -rf Phyloseq*_cache
# -- using R under base environment --
#(base) jhuang@WS-2290C:~/DATA_A/Data_Nicole8_Lamprecht_new_PUBLISHED/core_diversity_e1300
#mkdir figures
rmarkdown::render('Phyloseq.Rmd',output_file='Phyloseq.html')
2, bridges other tools
library(MicrobiotaProcess)
library(microeco)
library(ggalluvial)
library(ggh4x)
library(gghalves)
library(tidyr)
ps.ng.tax_sel <- ps.ng.tax_abund
#Choose all samples
otu_table(ps.ng.tax_sel) <- otu_table(ps.ng.tax_abund)[,c("kg001", "kg002", "kg003", "kg004", "kg005", "kg007", "kg009", "kg015", "kg016", "kg019", "kg020", "kg021", "kg022", "kg023", "kg025", "kg026", "kg027", "kg028", "kg029", "ra002", "ra003", "ra004", "ra005", "ra006", "ra007", "ra008", "ra009", "ra010", "ra013", "ra014", "ra015", "ra017", "ra018", "ra019", "ra020", "ra022", "ra023", "ra024", "ra025")]
mpse_abund <- ps.ng.tax_sel %>% as.MPSE()
3, rarefaction analysis
mpse_abund %<>% mp_rrarefy()
mpse_abund %<>%
mp_cal_rarecurve(
.abundance = RareAbundance,
chunks = 400
)
p1 <- mpse_abund %>%
mp_plot_rarecurve(
.rare = RareAbundanceRarecurve,
.alpha = Observe,
)
p2 <- mpse_abund %>%
mp_plot_rarecurve(
.rare = RareAbundanceRarecurve,
.alpha = Observe,
.group = SampleType
) +
scale_color_manual(values=c("#1f78b4", "#e31a1c")) +
scale_fill_manual(values=c("#1f78b4", "#e31a1c"), guide="none")
glimpse(mpse_abund)
mpse_abund %>% print(width=380, n=2)
p3 <- mpse_abund %>%
mp_plot_rarecurve(
.rare = RareAbundanceRarecurve,
.alpha = "Observe",
.group = SampleType,
plot.group = TRUE
) +
scale_color_manual(values=c("#1f78b4", "#e31a1c")) +
scale_fill_manual(values=c("#1f78b4", "#e31a1c"),guide="none")
png("rarefaction_of_samples_or_groups.png", width=1080, height=600)
p1 + p2 + p3
dev.off()
alpha diversity analysis
4, calculate alpha index and visualization
library(ggplot2)
library(MicrobiotaProcess)
mpse_abund %<>%
mp_cal_alpha(.abundance=RareAbundance)
mpse_abund
#NOTE mpse_abund contains 28 varibles = 22 varibles + Observe <dbl>, Chao1 <dbl>, ACE <dbl>, Shannon <dbl>, Simpson <dbl>, Pielou <dbl>
f1 <- mpse_abund %>%
mp_plot_alpha(
.group=SampleType,
.alpha=c(Observe, Chao1, ACE, Shannon, Simpson, Pielou)
) +
scale_fill_manual(values=c("#1f78b4", "#e31a1c"), guide="none") +
scale_color_manual(values=c("#1f78b4", "#e31a1c"), guide="none")
f2 <- mpse_abund %>%
mp_plot_alpha(
.alpha=c(Observe, Chao1, ACE, Shannon, Simpson, Pielou)
)
#ps.ng.tax_sel contais only pre samples --> f1 cannot be generated!
png("alpha_diversity_comparison.png", width=1400, height=600)
f1 / f2
dev.off()
5, visualize taxonomy abundance (Class)
mpse_abund %<>%
mp_cal_abundance( # for each samples
.abundance = RareAbundance
) %>%
mp_cal_abundance( # for each groups
.abundance=RareAbundance,
.group=SampleType
)
mpse_abund
p1 <- mpse_abund %>%
mp_plot_abundance(
.abundance=RareAbundance,
taxa.class = Class,
topn = 20,
relative = TRUE
)
p2 <- mpse_abund %>%
mp_plot_abundance(
.abundance=RareAbundance,
taxa.class = Class,
topn = 20,
relative = FALSE
)
png("relative_abundance_and_abundance.png", width= 1200, height=600) #NOT PRODUCED!
p1 / p2
dev.off()
h1 <- mpse_abund %>%
mp_plot_abundance(
.abundance = RareAbundance,
.group = SampleType,
taxa.class = Class,
relative = TRUE,
topn = 20,
geom = 'heatmap',
features.dist = 'euclidean',
features.hclust = 'average',
sample.dist = 'bray',
sample.hclust = 'average'
)
h2 <- mpse_abund %>%
mp_plot_abundance(
.abundance = RareAbundance,
.group = SampleType,
taxa.class = Class,
relative = FALSE,
topn = 20,
geom = 'heatmap',
features.dist = 'euclidean',
features.hclust = 'average',
sample.dist = 'bray',
sample.hclust = 'average'
)
# the character (scale or theme) of figure can be adjusted by set_scale_theme
# refer to the mp_plot_dist
png("relative_abundance_and_abundance_heatmap.png", width= 1200, height=600)
aplot::plot_list(gglist=list(h1, h2), tag_levels="A")
dev.off()
# visualize the relative abundance of top 20 class for each .group (SampleType)
p3 <- mpse_abund %>%
mp_plot_abundance(
.abundance=RareAbundance,
.group=SampleType,
taxa.class = Class,
topn = 20,
plot.group = TRUE
)
# visualize the abundance of top 20 phyla for each .group (time)
p4 <- mpse_abund %>%
mp_plot_abundance(
.abundance=RareAbundance,
.group= SampleType,
taxa.class = Class,
topn = 20,
relative = FALSE,
plot.group = TRUE
)
png("relative_abundance_and_abundance_groups.png", width= 1000, height=1000)
p3 / p4
dev.off()
beta diversity analysis
6, calculate the distance between samples or groups
mpse_abund %<>%
mp_decostand(.abundance=Abundance)
mpse_abund %<>% mp_cal_dist(.abundance=hellinger, distmethod="bray")
mpse_abund
p1 <- mpse_abund %>% mp_plot_dist(.distmethod = bray)
png("distance_between_samples.png", width= 1000, height=1000)
p1
dev.off()
# when .group is provided, the dot heatmap plot with group information will be return.
p2 <- mpse_abund %>% mp_plot_dist(.distmethod = bray, .group = SampleType)
# The scale or theme of dot heatmap plot can be adjusted using set_scale_theme function.
p2 %>% set_scale_theme(
x = scale_fill_manual(
values=c("#1f78b4", "#e31a1c"), #c("orange", "deepskyblue"),
guide = guide_legend(
keywidth = 1,
keyheight = 0.5,
title.theme = element_text(size=8),
label.theme = element_text(size=6)
)
),
aes_var = SampleType # specific the name of variable
) %>%
set_scale_theme(
x = scale_color_gradient(
guide = guide_legend(keywidth = 0.5, keyheight = 0.5)
),
aes_var = bray
) %>%
set_scale_theme(
x = scale_size_continuous(
range = c(0.1, 3),
guide = guide_legend(keywidth = 0.5, keyheight = 0.5)
),
aes_var = bray
)
png("distance_between_samples_with_group_info.png", width= 1000, height=1000)
p2
dev.off()
# when .group is provided and group.test is TRUE, the comparison of different groups will be returned
# Assuming p3 is a ggplot object after mp_plot_dist call
p3 <- mpse_abund %>%
mp_plot_dist(.distmethod = bray, .group = SampleType, group.test = TRUE, textsize = 6) +
theme(
axis.title.x = element_text(size = 14), # Customize x-axis label face = "bold"
axis.title.y = element_text(size = 14), # Customize y-axis label
axis.text.x = element_text(size = 14), # Customize x-axis ticks
axis.text.y = element_text(size = 14) # Customize y-axis ticks
)
# Save the plot with the new theme settings
png("Comparison_of_Bray_Distances.png", width = 1000, height = 1000)
print(p3) # Ensure that p3 is explicitly printed in the device
dev.off()
# Extract Bray-Curtis Distance Values and save them in a Excel-table.
library(dplyr)
library(openxlsx)
# Define the sample numbers vector
sample_numbers <- c("1","2","5","6","7", "29","30","31","32")
# Consolidate the list of tibbles using the actual sample numbers
bray_data <- bind_rows(
lapply(seq_along(mpse_abund$bray), function(i) {
tibble(
Sample1 = sample_numbers[i], # Use actual sample number
Sample2 = mpse_abund$bray[[i]]$braySampley,
BrayDistance = mpse_abund$bray[[i]]$bray
)
}),
.id = "PairID"
)
# Print the data frame to check the output
print(bray_data)
# Write the data frame to an Excel file
write.xlsx(bray_data, file = "Bray_Curtis_Distances.xlsx")
#DELETE the column "PairID" in Excel file
7, the PCoA analysis
#install.packages("corrr")
library(corrr)
#install.packages("ggside")
library(ggside)
mpse_abund %<>%
mp_cal_pcoa(.abundance=hellinger, distmethod="bray")
# The dimensions of ordination analysis will be added the colData slot (default).
mpse_abund
methods(class=class(mpse_abund))
mpse_abund %>% print(width=380, n=2)
#NOTE mpse_abund contains 34 varibles = 31 varibles + `PCo1 (30.16%)` <dbl>, `PCo2 (15.75%)` <dbl>, `PCo3 (10.53%)` <dbl> + [Domain ... Species]
# We also can perform adonis or anosim to check whether it is significant to the dissimilarities of groups.
mpse_abund %<>%
mp_adonis(.abundance=hellinger, .formula=~SampleType, distmethod="bray", permutations=9999, action="add")
mpse_abund %>% mp_extract_internal_attr(name=adonis)
#PAUSE
p1 <- mpse_abund %>%
mp_plot_ord(
.ord = pcoa,
.group = SampleType,
.color = SampleType,
.size = 2.4,
.alpha = 1,
ellipse = TRUE,
show.legend = FALSE # don't display the legend of stat_ellipse
) +
scale_fill_manual(
#values = c("#a6cee3", "#1f78b4", "#b2df8a", "#33a02c", "#fb9a99", "#e31a1c", "#cab2d6", "#6a3d9a"),
#values = c("#a6cee3", "#b2df8a", "#fb9a99", "#cab2d6"),
values = c("#1f78b4", "#e31a1c"),
guide = guide_legend(keywidth=1.6, keyheight=1.6, label.theme=element_text(size=12))
) +
scale_color_manual(
#values=c("#a6cee3", "#1f78b4", "#b2df8a", "#33a02c", "#fb9a99", "#e31a1c", "#cab2d6", "#6a3d9a"),
#values = c("#a6cee3", "#b2df8a", "#fb9a99", "#cab2d6"),
values = c("#1f78b4", "#e31a1c"),
guide = guide_legend(keywidth=1.6, keyheight=1.6, label.theme=element_text(size=12))
)
pdf("PCoA.pdf")
p1
dev.off()
# The size of point also can be mapped to other variables such as Observe, or Shannon
# Then the alpha diversity and beta diversity will be displayed simultaneously.
p2 <- mpse_abund %>%
mp_plot_ord(
.ord = pcoa,
.group = SampleType,
.color = SampleType,
.size = Shannon,
.alpha = Observe,
ellipse = TRUE,
show.legend = FALSE # don't display the legend of stat_ellipse
) +
scale_fill_manual(
values = c("#1f78b4", "#e31a1c"), #only needs four colors.
#values = c("#FF0000", "#000000", "#0000FF", "#C0C0C0", "#00FF00", "#FFFF00", "#00FFFF", "#FFA500"),
guide = guide_legend(keywidth=0.6, keyheight=0.6, label.theme=element_text(size=8))
) +
scale_color_manual(
values = c("#1f78b4", "#e31a1c"), #only needs four colors.
#values=c("#FF0000", "#000000", "#0000FF", "#C0C0C0", "#00FF00", "#FFFF00", "#00FFFF", "#FFA500"),
guide = guide_legend(keywidth=0.6, keyheight=0.6, label.theme=element_text(size=8))
) +
scale_size_continuous(
range=c(0.5, 3),
guide = guide_legend(keywidth=0.6, keyheight=0.6, label.theme=element_text(size=8))
)
pdf("PCoA2.pdf")
p2
dev.off()
# Add the sample name as text labels
library(ggrepel)
p2 <- mpse_abund %>%
mp_plot_ord(
.ord = pcoa,
.group = SampleType,
.color = SampleType,
.size = Shannon,
.alpha = Observe,
ellipse = TRUE,
show.legend = FALSE # don't display the legend of stat_ellipse
) +
geom_text_repel(aes(label = ifelse(Sample == "1", "1", Sample)), # Prioritize "1"
size = 3,
color = "black", # Set the label color to black for better visibility
max.overlaps = Inf, # Allow maximum labels
force = 2, # Increase the force to push labels apart
box.padding = 0.5, # Add more padding around the labels
segment.size = 0.2 # Line segment size connecting labels to points
) +
scale_fill_manual(
values = c("#1f78b4", "#e31a1c"), # only needs two colors
guide = guide_legend(keywidth=0.6, keyheight=0.6, label.theme=element_text(size=8))
) +
scale_color_manual(
values = c("#1f78b4", "#e31a1c"), # only needs two colors
guide = guide_legend(keywidth=0.6, keyheight=0.6, label.theme=element_text(size=8))
) +
scale_size_continuous(
range=c(0.5, 3),
guide = guide_legend(keywidth=0.6, keyheight=0.6, label.theme=element_text(size=8))
)
#pdf("PCoA2_labeled.pdf")
png("PCoA2_labeled.png", width=800, height=800)
p2
dev.off()
8, hierarchical cluster (tree) analysis
#input should contain hellinger!
mpse_abund %<>%
mp_cal_clust(
.abundance = hellinger,
distmethod = "bray",
hclustmethod = "average", # (UPGAE)
action = "add" # action is used to control which result will be returned
)
mpse_abund
mpse_abund %>% print(width=380, n=2)
#NOTE mpse_abund contains 34 varibles, no new variable, the column bray has been new calculated!
# if action = 'add', the result of hierarchical cluster will be added to the MPSE object
# mp_extract_internal_attr can extract it. It is a treedata object, so it can be visualized
# by ggtree.
sample.clust <- mpse_abund %>% mp_extract_internal_attr(name='SampleClust')
#The object contained internal attribute: PCoA ADONIS SampleClust
sample.clust
#--> The associated data tibble abstraction: 27 × 30
library(ggtree)
p <- ggtree(sample.clust) +
geom_tippoint(aes(color=SampleType)) +
geom_tiplab(as_ylab = TRUE) +
ggplot2::scale_x_continuous(expand=c(0, 0.01))
png("hierarchical_cluster1.png", width= 1000, height=800)
p
dev.off()
#https://bioconductor.org/packages/release/bioc/vignettes/MicrobiotaProcess/inst/doc//MicrobiotaProcess.html
# mapping = aes(x = RelRareAbundanceBySample-->SampleType,
# y = Sample-->SampleType,
# fill = Phyla
# ),
library(ggtreeExtra)
library(ggplot2)
# Extract relative abundance of phyla
phyla.tb <- mpse_abund %>%
mp_extract_abundance(taxa.class=Phylum, topn=30)
# The abundance of each samples is nested, it can be flatted using the unnest of tidyr.
phyla.tb %<>% tidyr::unnest(cols=RareAbundanceBySample) %>% dplyr::rename(Phyla="label")
phyla.tb
phyla.tb %>% print(width=380, n=10)
p1 <- p +
geom_fruit(
data=phyla.tb,
geom=geom_col,
mapping = aes(x = RelRareAbundanceBySample,
y = Sample,
fill = Phyla
),
orientation = "y",
#offset = 0.4,
pwidth = 3,
axis.params = list(axis = "x",
title = "The relative abundance of phyla (%)",
title.size = 4,
text.size = 2,
vjust = 1),
grid.params = list()
)
png("hierarchical_cluster2_Phyla.png", width = 1000, height = 800)
p1
dev.off()
# Extract relative abundance of classes
class.tb <- mpse_abund %>%
mp_extract_abundance(taxa.class = Class, topn = 30)
# Flatten and rename the columns
class.tb %<>% tidyr::unnest(cols = RareAbundanceBySample) %>% dplyr::rename(Class = "label")
# View the data frame
class.tb
# Create the plot
p1 <- p +
geom_fruit(
data = class.tb,
geom = geom_col,
mapping = aes(x = RelRareAbundanceBySample,
y = Sample,
fill = Class
),
orientation = "y",
pwidth = 3,
axis.params = list(axis = "x",
title = "The relative abundance of classes (%)",
title.size = 4,
text.size = 2,
vjust = 1),
grid.params = list()
)
# Save the plot to a file #ERROR-->NEED to be DEBUGGED!
png("hierarchical_cluster2_Class.png", width = 1000, height = 800)
print(p1)
dev.off()
9, biomarker discovery (update Sign_Group to Sign_SampleType, RareAbundanceByGroup to RareAbundanceBySampleType)
library(ggtree)
library(ggtreeExtra)
library(ggplot2)
library(MicrobiotaProcess)
library(tidytree)
library(ggstar)
library(forcats)
library(writexl)
mpse_abund %>% print(width=150)
#mpse_abund %<>%
# mp_cal_abundance( # for each samples
# .abundance = RareAbundance
# ) %>%
# mp_cal_abundance( # for each groups
# .abundance=RareAbundance,
# .group=SampleType
# )
#mpse_abund
mpse_abund %<>%
mp_diff_analysis(
.abundance = RelRareAbundanceBySample,
.group = SampleType,
cl.min = 4,
first.test.alpha = 0.01, filter.p="pvalue"
)
# The result is stored to the taxatree or otutree slot, you can use mp_extract_tree to extract the specific slot.
taxa.tree <- mpse_abund %>%
mp_extract_tree(type="taxatree")
taxa.tree
## And the result tibble of different analysis can also be extracted with tidytree (>=0.3.5)
#LDAupper, LDAmean, LDAlower,
taxa.tree %>% select(label, nodeClass, Sign_SampleType, fdr) #%>% dplyr::filter(!is.na(fdr))
taxa.tree %>% print(width=150, n=200)
# -- replace the pvalue and fdr with pvalue and p-adjusted from DESeq enrichment results --
tree_data <- as_tibble(taxa.tree)
# ---- modify tree_data by left_joining with sigtab and updating Sign_SampleType ----
sigtab$label <- rownames(sigtab)
write.xlsx(sigtab, file = "sigtab.xlsx")
sum(sigtab$padj<0.05)
#taxa.tree <- left_join(tree_data, sigtab[, c("label", "log2FoldChange", "pvalue", "padj")], by = 'label') %>% as.treedata
taxa.tree2 <- tree_data %>%
left_join(sigtab[, c("label", "baseMean", "log2FoldChange", "lfcSE", "stat", "pvalue", "padj")], by = "label") %>%
mutate(Sign_SampleType = case_when(
log2FoldChange > 0 & padj <= 0.05 ~ "RA",
log2FoldChange < 0 & padj <= 0.05 ~ "control",
TRUE ~ NA_character_ # Sets Sign_SampleType to NA otherwise
)) %>%
as.treedata() # Convert the dataframe to a treedata object
# ---- print taxa_data2 to Excel ----
taxa.tree2 %>% print(width=380, n=20)
taxa_data2 <- as_tibble(taxa.tree2)
sum(!is.na(taxa_data2$Sign_SampleType))
sapply(taxa_data2, class)
# Remove or transform list columns if not needed
taxa_data2_simplified <- taxa_data2 %>%
select(-RareAbundanceBySample, -RareAbundanceBySampleType) %>%
mutate(across(where(is.list), ~toString(.))) # Convert lists to character strings if needed
# Replace NA with a placeholder, such as "NA" or another suitable representation
taxa_data2_simplified <- taxa_data2_simplified %>%
mutate(across(everything(), ~ifelse(is.na(.), "NA", .)))
taxonomy_data <- as.data.frame(mp_extract_taxonomy(mpse_abund))
colnames(taxa_data2_simplified)[colnames(taxa_data2_simplified) == "label"] <- "OTU"
combined_data <- left_join(taxa_data2_simplified, taxonomy_data, by = "OTU")
write_xlsx(combined_data, "taxa_data2.xlsx")
#(UNDER HOST-ENV) cp sigtab.xlsx diff_analysis_RA_vs_control.xlsx and then switch label as the 1st column and sort the columns by padj.
# -- NOTE that sometimes the record in DESeq2 not occurs in the final list, since the statistics calculation of MicrobiotaProcess results in NA, e.g. the record FJ879443.1.1488, we can simply delete the record from diff_analysis_RA_vs_control.xlsx --
# ---- since taxa.tree is treedata object, it can be visualized by ggtree and ggtreeExtra ----
p1 <- ggtree(
taxa.tree2,
layout="radial",
size = 0.3
) +
geom_point(
data = td_filter(!isTip),
fill="white",
size=1,
shape=21
)
# display the high light of phylum clade.
p2 <- p1 +
geom_hilight(
data = td_filter(nodeClass == "Phylum"),
mapping = aes(node = node, fill = label)
)
# display the relative abundance of features(OTU)
p3 <- p2 +
ggnewscale::new_scale("fill") +
geom_fruit(
data = td_unnest(RareAbundanceBySample),
geom = geom_star,
mapping = aes(
x = fct_reorder(Sample, SampleType, .fun=min),
size = RelRareAbundanceBySample,
fill = SampleType,
subset = RelRareAbundanceBySample > 0
),
starshape = 13,
starstroke = 0.25,
offset = 0.03,
pwidth = 0.4,
grid.params = list(linetype=2)
) +
scale_size_continuous(
name="Relative Abundance (%)",
range = c(.5, 3)
) +
scale_fill_manual(values=c("#1B9E77", "#D95F02"))
# display the tip labels of taxa tree
p4 <- p3 + geom_tiplab(size=6, offset=4.0)
# display the LDA of significant OTU.
#p5 <- p4 +
# ggnewscale::new_scale("fill") +
# geom_fruit(
# geom = geom_col,
# mapping = aes(
# x = LDAmean,
# fill = Sign_SampleType,
# subset = !is.na(LDAmean)
# ),
# orientation = "y",
# offset = 0.3,
# pwidth = 0.5,
# axis.params = list(axis = "x",
# title = "Log10(LDA)",
# title.height = 0.01,
# title.size = 2,
# text.size = 1.8,
# vjust = 1),
# grid.params = list(linetype = 2)
# )
# display the significant (FDR-->pvalue-->padj) taxonomy after kruskal.test (default)
#shape = 21,
#scale_size_continuous(range=c(1, 3)) +
p6 <- p4 +
ggnewscale::new_scale("size") +
geom_point(
data=td_filter(!is.na(Sign_SampleType)),
mapping = aes(size = -log10(padj),
fill = Sign_SampleType,
),
shape = 21,
) +
scale_size_continuous(range=c(1, 4)) +
scale_fill_manual(values=c("#1B9E77", "#D95F02"))
svg("diff_analysis.svg",width=22, height=22)
#png("differently_expressed_otu.png", width=2000, height=2000)
p6 + theme(
legend.key.height = unit(1.0, "cm"),
legend.key.width = unit(1.0, "cm"),
legend.spacing.y = unit(0.01, "cm"),
legend.text = element_text(size = 20),
legend.title = element_text(size = 20)
#legend.position = c(0.99, 0.01)
)
dev.off()
点赞本文的读者
还没有人对此文章表态
本文有评论
没有评论
看文章,发评论,不要沉默
最受欢迎文章
- Motif Discovery in Biological Sequences: A Comparison of MEME and HOMER
- Why Do Significant Gene Lists Change After Adding Additional Conditions in Differential Gene Expression Analysis?
- Calling peaks using findPeaks of HOMER
- PiCRUST2 Pipeline for Functional Prediction and Pathway Analysis in Metagenomics
- Updating Human Gene Identifiers using Ensembl BioMart: A Step-by-Step Guide
- pheatmap vs heatmap.2
- Should the inputs for GSVA be normalized or raw?
- Setup conda environments
- Kraken2 Installation and Usage Guide
- File format for single channel analysis of Agilent microarray data with Limma?
最新文章
- Setup the environment for lumicks-pylake and C_Trap-Multimer-photontrack.ipynb
- 🧬 Cadmium Resistance Gene Analysis in Staphylococcus epidermidis HD46
- MCV病毒中的LT与sT蛋白功能
- Analysis of the RNA binding protein (RBP) motifs for RNA-Seq and miRNAs (v3, simplied)
最多评论文章
- Updating Human Gene Identifiers using Ensembl BioMart: A Step-by-Step Guide
- The top 10 genes
- Retrieving KEGG Genes Using Bioservices in Python
推荐相似文章
Sorry, 没有相似文章